ADAR1 RNA editing regulates endothelial cell functions via the MDA-5 RNA sensing signaling pathway
- PMID:34969816
- PMCID: PMC8739526
- DOI: 10.26508/lsa.202101191
ADAR1 RNA editing regulates endothelial cell functions via the MDA-5 RNA sensing signaling pathway
Abstract
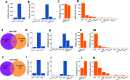
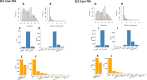
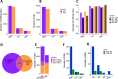
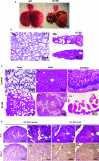
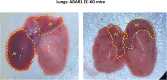
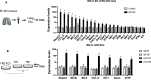
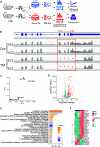
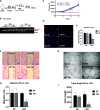
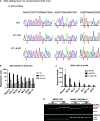
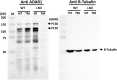
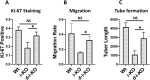
The RNA-sensing signaling pathway has been well studied as an essential antiviral mechanism of innate immunity. However, its role in non-infected cells is yet to be thoroughly characterized. Here, we demonstrated that the RNA sensing signaling pathway also reacts to the endogenous cellular RNAs in endothelial cells (ECs), and this reaction is regulated by the RNA-editing enzyme ADAR1. Cellular RNA sequencing analysis showed that EC RNAs endure extensive RNA editing, especially in the RNA transcripts of short interspersed nuclear elements. The EC-specific deletion of ADAR1 dramatically reduced the editing level on short interspersed nuclear element RNAs, resulting in newborn death in mice with damage evident in multiple organs. Genome-wide gene expression analysis revealed a prominent innate immune activation with a dramatically elevated expression of interferon-stimulated genes. However, blocking the RNA sensing signaling pathway by deletion of the cellular RNA receptor MDA-5 prevented interferon-stimulated gene expression and rescued the newborn mice from death. This evidence demonstrated that the RNA-editing/RNA-sensing signaling pathway dramatically modulates EC function, representing a novel molecular mechanism for the regulation of EC functions.
© 2021 Guo et al.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures










Similar articles
- The RNA-editing enzyme ADAR1: a regulatory hub that tunes multiple dsRNA-sensing pathways.Nakahama T, Kawahara Y.Nakahama T, et al.Int Immunol. 2023 Mar 14;35(3):123-133. doi: 10.1093/intimm/dxac056.Int Immunol. 2023.PMID:36469491
- RNA editing by ADAR1 prevents MDA5 sensing of endogenous dsRNA as nonself.Liddicoat BJ, Piskol R, Chalk AM, Ramaswami G, Higuchi M, Hartner JC, Li JB, Seeburg PH, Walkley CR.Liddicoat BJ, et al.Science. 2015 Sep 4;349(6252):1115-20. doi: 10.1126/science.aac7049. Epub 2015 Jul 23.Science. 2015.PMID:26275108Free PMC article.
- ADAR1 Prevents Autoinflammatory Processes in the Heart Mediated by IRF7.Garcia-Gonzalez C, Dieterich C, Maroli G, Wiesnet M, Wietelmann A, Li X, Yuan X, Graumann J, Stellos K, Kubin T, Schneider A, Braun T.Garcia-Gonzalez C, et al.Circ Res. 2022 Sep 16;131(7):580-597. doi: 10.1161/CIRCRESAHA.122.320839. Epub 2022 Aug 24.Circ Res. 2022.PMID:36000401
- The role of RNA editing by ADAR1 in prevention of innate immune sensing of self-RNA.Heraud-Farlow JE, Walkley CR.Heraud-Farlow JE, et al.J Mol Med (Berl). 2016 Oct;94(10):1095-1102. doi: 10.1007/s00109-016-1416-1. Epub 2016 Apr 5.J Mol Med (Berl). 2016.PMID:27044320Review.
- What do editors do? Understanding the physiological functions of A-to-I RNA editing by adenosine deaminase acting on RNAs.Heraud-Farlow JE, Walkley CR.Heraud-Farlow JE, et al.Open Biol. 2020 Jul;10(7):200085. doi: 10.1098/rsob.200085. Epub 2020 Jul 1.Open Biol. 2020.PMID:32603639Free PMC article.Review.
Cited by
- ADAR1 suppression causes interferon signaling and transposable element transcript accumulation in human astrocytes.McEntee CM, Cavalier AN, LaRocca TJ.McEntee CM, et al.Front Mol Neurosci. 2023 Oct 25;16:1263369. doi: 10.3389/fnmol.2023.1263369. eCollection 2023.Front Mol Neurosci. 2023.PMID:38035265Free PMC article.
- RNA modifications in cardiovascular health and disease.Gatsiou A, Stellos K.Gatsiou A, et al.Nat Rev Cardiol. 2023 May;20(5):325-346. doi: 10.1038/s41569-022-00804-8. Epub 2022 Dec 5.Nat Rev Cardiol. 2023.PMID:36471164Review.
- Functional roles of epitranscriptomic marks in the cardiovascular system and disease: a narrative review.Ilieva M, Uchida S.Ilieva M, et al.Ann Transl Med. 2022 Jul;10(13):753. doi: 10.21037/atm-22-1074.Ann Transl Med. 2022.PMID:35957723Free PMC article.Review.
- Case Report: Aicardi-Goutières Syndrome Type 6 and Dyschromatosis Symmetrica Hereditaria With Congenital Heart Disease and Mitral Valve Calcification - Phenotypic Variants Caused by Adenosine Deaminase Acting on the RNA 1 Gene Homozygous Mutations.Liu L, Zhang L, Huang P, Xiong J, Xiao Y, Wang C, Mao D, Liu L.Liu L, et al.Front Pediatr. 2022 Jun 27;10:852903. doi: 10.3389/fped.2022.852903. eCollection 2022.Front Pediatr. 2022.PMID:35832578Free PMC article.
- An AGS-associated mutation in ADAR1 catalytic domain results in early-onset and MDA5-dependent encephalopathy with IFN pathway activation in the brain.Guo X, Steinman RA, Sheng Y, Cao G, Wiley CA, Wang Q.Guo X, et al.J Neuroinflammation. 2022 Dec 1;19(1):285. doi: 10.1186/s12974-022-02646-0.J Neuroinflammation. 2022.PMID:36457126Free PMC article.
References
Publication types
MeSH terms
Substances
Related information
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases