Comparative host-coronavirus protein interaction networks reveal pan-viral disease mechanisms
- PMID:33060197
- PMCID: PMC7808408
- DOI: 10.1126/science.abe9403
Comparative host-coronavirus protein interaction networks reveal pan-viral disease mechanisms
Abstract
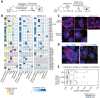
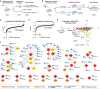
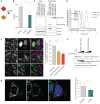
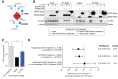
The COVID-19 pandemic, caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), is a grave threat to public health and the global economy. SARS-CoV-2 is closely related to the more lethal but less transmissible coronaviruses SARS-CoV-1 and Middle East respiratory syndrome coronavirus (MERS-CoV). Here, we have carried out comparative viral-human protein-protein interaction and viral protein localization analyses for all three viruses. Subsequent functional genetic screening identified host factors that functionally impinge on coronavirus proliferation, including Tom70, a mitochondrial chaperone protein that interacts with both SARS-CoV-1 and SARS-CoV-2 ORF9b, an interaction we structurally characterized using cryo-electron microscopy. Combining genetically validated host factors with both COVID-19 patient genetic data and medical billing records identified molecular mechanisms and potential drug treatments that merit further molecular and clinical study.
Copyright © 2020 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Figures






Comment in
- Conserved host-pathogen interactions identify novel treatment options in betacoronavirus infections.Lukassen S, Eils R.Lukassen S, et al.Signal Transduct Target Ther. 2021 Feb 10;6(1):57. doi: 10.1038/s41392-021-00480-z.Signal Transduct Target Ther. 2021.PMID:33563888Free PMC article.No abstract available.
- Olanzapine, risperidone and quetiapine: Do these atypical antipsychotics have a protective effect for SARS-CoV-2?Prokopez CR, Farinola R, Vallejos M, Lopredo LS, Sfriso LE, Chiapella LC, Arce C, Corral RM, Cuesta MJ, Alomo M.Prokopez CR, et al.Schizophr Res. 2022 Mar;241:140-141. doi: 10.1016/j.schres.2022.01.035. Epub 2022 Jan 24.Schizophr Res. 2022.PMID:35123336Free PMC article.No abstract available.
Similar articles
- Crystal structure of SARS-CoV-2 Orf9b in complex with human TOM70 suggests unusual virus-host interactions.Gao X, Zhu K, Qin B, Olieric V, Wang M, Cui S.Gao X, et al.Nat Commun. 2021 May 14;12(1):2843. doi: 10.1038/s41467-021-23118-8.Nat Commun. 2021.PMID:33990585Free PMC article.
- The Nucleocapsid (N) Proteins of Different Human Coronaviruses Demonstrate a Variable Capacity to Induce the Formation of Cytoplasmic Condensates.Tikhomirova MA, Kuzmenko OL, Arifulin EA, Shirokova OM, Musinova YR, Sheval EV.Tikhomirova MA, et al.Int J Mol Sci. 2024 Dec 7;25(23):13162. doi: 10.3390/ijms252313162.Int J Mol Sci. 2024.PMID:39684875Free PMC article.
- Dysregulation of lung epithelial cell homeostasis and immunity contributes to Middle East respiratory syndrome coronavirus disease severity.Sims AC, Schäfer A, Okuda K, Leist SR, Kocher JF, Cockrell AS, Hawkins PE, Furusho M, Jensen KL, Kyle JE, Burnum-Johnson KE, Stratton KG, Lamar NC, Niccora CD, Weitz KK, Smith RD, Metz TO, Waters KM, Boucher RC, Montgomery SA, Baric RS, Sheahan TP.Sims AC, et al.mSphere. 2025 Feb 25;10(2):e0095124. doi: 10.1128/msphere.00951-24. Epub 2025 Jan 30.mSphere. 2025.PMID:39882872Free PMC article.
- Genetic variants of the human host influencing the coronavirus-associated phenotypes (SARS, MERS and COVID-19): rapid systematic review and field synopsis.Di Maria E, Latini A, Borgiani P, Novelli G.Di Maria E, et al.Hum Genomics. 2020 Sep 11;14(1):30. doi: 10.1186/s40246-020-00280-6.Hum Genomics. 2020.PMID:32917282Free PMC article.Review.
- Antibody tests for identification of current and past infection with SARS-CoV-2.Fox T, Geppert J, Dinnes J, Scandrett K, Bigio J, Sulis G, Hettiarachchi D, Mathangasinghe Y, Weeratunga P, Wickramasinghe D, Bergman H, Buckley BS, Probyn K, Sguassero Y, Davenport C, Cunningham J, Dittrich S, Emperador D, Hooft L, Leeflang MM, McInnes MD, Spijker R, Struyf T, Van den Bruel A, Verbakel JY, Takwoingi Y, Taylor-Phillips S, Deeks JJ; Cochrane COVID-19 Diagnostic Test Accuracy Group.Fox T, et al.Cochrane Database Syst Rev. 2022 Nov 17;11(11):CD013652. doi: 10.1002/14651858.CD013652.pub2.Cochrane Database Syst Rev. 2022.PMID:36394900Free PMC article.Review.
Cited by
- Molecular Bases of Serotonin Reuptake Inhibitor Antidepressant-Attributed Effects in COVID-19: A New Insight on the Role of Bradykinins.Gouda AS, Mégarbane B.Gouda AS, et al.J Pers Med. 2022 Sep 11;12(9):1487. doi: 10.3390/jpm12091487.J Pers Med. 2022.PMID:36143272Free PMC article.Review.
- Recent computational drug repositioning strategies against SARS-CoV-2.Lu L, Qin J, Chen J, Yu N, Miyano S, Deng Z, Li C.Lu L, et al.Comput Struct Biotechnol J. 2022;20:5713-5728. doi: 10.1016/j.csbj.2022.10.017. Epub 2022 Oct 17.Comput Struct Biotechnol J. 2022.PMID:36277237Free PMC article.
- Navigating the landscape of mitochondrial-ER communication in health and disease.Ronayne CT, Latorre-Muro P.Ronayne CT, et al.Front Mol Biosci. 2024 Jan 23;11:1356500. doi: 10.3389/fmolb.2024.1356500. eCollection 2024.Front Mol Biosci. 2024.PMID:38323074Free PMC article.Review.
- A comprehensive library of fluorescent constructs of SARS-CoV-2 proteins and their initial characterisation in different cell types.Miserey-Lenkei S, Trajkovic K, D'Ambrosio JM, Patel AJ, Čopič A, Mathur P, Schauer K, Goud B, Albanèse V, Gautier R, Subra M, Kovacs D, Barelli H, Antonny B.Miserey-Lenkei S, et al.Biol Cell. 2021 Jul;113(7):311-328. doi: 10.1111/boc.202000158. Epub 2021 May 10.Biol Cell. 2021.PMID:33666950Free PMC article.
- A Cullin 5-based complex serves as an essential modulator of ORF9b stability in SARS-CoV-2 replication.Zhou Y, Chen Z, Liu S, Liu S, Liao Y, Du A, Dong Z, Zhang Y, Chen X, Tao S, Wu X, Razzaq A, Xu G, Tan DA, Li S, Deng Y, Peng J, Dai S, Deng X, Zhang X, Jiang T, Zhang Z, Cheng G, Zhao J, Xia Z.Zhou Y, et al.Signal Transduct Target Ther. 2024 Jun 28;9(1):159. doi: 10.1038/s41392-024-01874-5.Signal Transduct Target Ther. 2024.PMID:38937432Free PMC article.
References
- Beigel J. H., Tomashek K. M., Dodd L. E., Mehta A. K., Zingman B. S., Kalil A. C., Hohmann E., Chu H. Y., Luetkemeyer A., Kline S., Lopez de Castilla D., Finberg R. W., Dierberg K., Tapson V., Hsieh L., Patterson T. F., Paredes R., Sweeney D. A., Short W. R., Touloumi G., Lye D. C., Ohmagari N., Oh M.-d., Ruiz-Palacios G. M., Benfield T., Fätkenheuer G., Kortepeter M. G., Atmar R. L., Creech C. B., Lundgren J., Babiker A. G., Pett S., Neaton J. D., Burgess T. H., Bonnett T., Green M., Makowski M., Osinusi A., Nayak S., Lane H. C., ACTT-1 Study Group Members , Remdesivir for the treatment of Covid-19—Final report. N. Engl. J. Med. 383, 1813–1826 (2020). 10.1056/NEJMoa2007764 - DOI - PMC - PubMed
- Gordon D. E., Jang G. M., Bouhaddou M., Xu J., Obernier K., White K. M., O’Meara M. J., Rezelj V. V., Guo J. Z., Swaney D. L., Tummino T. A., Hüttenhain R., Kaake R. M., Richards A. L., Tutuncuoglu B., Foussard H., Batra J., Haas K., Modak M., Kim M., Haas P., Polacco B. J., Braberg H., Fabius J. M., Eckhardt M., Soucheray M., Bennett M. J., Cakir M., McGregor M. J., Li Q., Meyer B., Roesch F., Vallet T., Mac Kain A., Miorin L., Moreno E., Naing Z. Z. C., Zhou Y., Peng S., Shi Y., Zhang Z., Shen W., Kirby I. T., Melnyk J. E., Chorba J. S., Lou K., Dai S. A., Barrio-Hernandez I., Memon D., Hernandez-Armenta C., Lyu J., Mathy C. J. P., Perica T., Pilla K. B., Ganesan S. J., Saltzberg D. J., Rakesh R., Liu X., Rosenthal S. B., Calviello L., Venkataramanan S., Liboy-Lugo J., Lin Y., Huang X.-P., Liu Y., Wankowicz S. A., Bohn M., Safari M., Ugur F. S., Koh C., Savar N. S., Tran Q. D., Shengjuler D., Fletcher S. J., O’Neal M. C., Cai Y., Chang J. C. J., Broadhurst D. J., Klippsten S., Sharp P. P., Wenzell N. A., Kuzuoglu-Ozturk D., Wang H.-Y., Trenker R., Young J. M., Cavero D. A., Hiatt J., Roth T. L., Rathore U., Subramanian A., Noack J., Hubert M., Stroud R. M., Frankel A. D., Rosenberg O. S., Verba K. A., Agard D. A., Ott M., Emerman M., Jura N., von Zastrow M., Verdin E., Ashworth A., Schwartz O., d’Enfert C., Mukherjee S., Jacobson M., Malik H. S., Fujimori D. G., Ideker T., Craik C. S., Floor S. N., Fraser J. S., Gross J. D., Sali A., Roth B. L., Ruggero D., Taunton J., Kortemme T., Beltrao P., Vignuzzi M., García-Sastre A., Shokat K. M., Shoichet B. K., Krogan N. J., A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583, 459–468 (2020). 10.1038/s41586-020-2286-9 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Related information
Grants and funding
- T32 AI060537/AI/NIAID NIH HHS/United States
- HHSN272201400008C/AI/NIAID NIH HHS/United States
- T32 GM007618/GM/NIGMS NIH HHS/United States
- R01 AI122747/AI/NIAID NIH HHS/United States
- F32 CA239333/CA/NCI NIH HHS/United States
- R01 AI128214/AI/NIAID NIH HHS/United States
- R35 GM122481/GM/NIGMS NIH HHS/United States
- P01 AI063302/AI/NIAID NIH HHS/United States
- P50 AI150476/AI/NIAID NIH HHS/United States
- MC_UU_12016/2/MRC_/Medical Research Council/United Kingdom
- BB/J014443/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- R01 GM024485/GM/NIGMS NIH HHS/United States
- F30 AI143401/AI/NIAID NIH HHS/United States
- R01 AI120694/AI/NIAID NIH HHS/United States
- T32 GM008284/GM/NIGMS NIH HHS/United States
- MC_PC_19026/MRC_/Medical Research Council/United Kingdom
- R01 HG008742/HG/NHGRI NIH HHS/United States
- HHMI/Howard Hughes Medical Institute/United States
- U19 AI135990/AI/NIAID NIH HHS/United States
- F32 GM137463/GM/NIGMS NIH HHS/United States
- 201366/Z/16/Z/WT_/Wellcome Trust/United Kingdom
- R01 NS089713/NS/NINDS NIH HHS/United States
- R35 GM118099/GM/NIGMS NIH HHS/United States
- R01 HG009979/HG/NHGRI NIH HHS/United States
- MR/V03541X/1/MRC_/Medical Research Council/United Kingdom
- MC_UU_00018/1/MRC_/Medical Research Council/United Kingdom
- P01 AI120943/AI/NIAID NIH HHS/United States
- U19 AI135972/AI/NIAID NIH HHS/United States
- MC_UU_12014/2/MRC_/Medical Research Council/United Kingdom
- K99 GM138753/GM/NIGMS NIH HHS/United States
- F32 CA239336/CA/NCI NIH HHS/United States
- R01 AI143292/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous
