Optogenetic determination of the myocardial requirements for extrasystoles by cell type-specific targeting of ChannelRhodopsin-2
- PMID:26204914
- PMCID: PMC4538656
- DOI: 10.1073/pnas.1509380112
Optogenetic determination of the myocardial requirements for extrasystoles by cell type-specific targeting of ChannelRhodopsin-2
Abstract
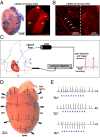
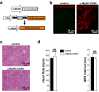
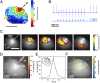
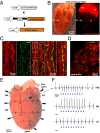
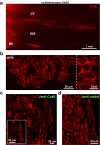
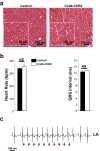
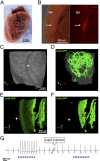
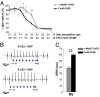
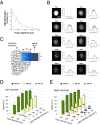
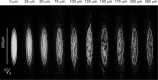
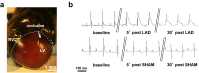
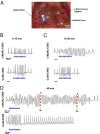
Extrasystoles lead to several consequences, ranging from uneventful palpitations to lethal ventricular arrhythmias, in the presence of pathologies, such as myocardial ischemia. The role of working versus conducting cardiomyocytes, as well as the tissue requirements (minimal cell number) for the generation of extrasystoles, and the properties leading ectopies to become arrhythmia triggers (topology), in the normal and diseased heart, have not been determined directly in vivo. Here, we used optogenetics in transgenic mice expressing ChannelRhodopsin-2 selectively in either cardiomyocytes or the conduction system to achieve cell type-specific, noninvasive control of heart activity with high spatial and temporal resolution. By combining measurement of optogenetic tissue activation in vivo and epicardial voltage mapping in Langendorff-perfused hearts, we demonstrated that focal ectopies require, in the normal mouse heart, the simultaneous depolarization of at least 1,300-1,800 working cardiomyocytes or 90-160 Purkinje fibers. The optogenetic assay identified specific areas in the heart that were highly susceptible to forming extrasystolic foci, and such properties were correlated to the local organization of the Purkinje fiber network, which was imaged in three dimensions using optical projection tomography. Interestingly, during the acute phase of myocardial ischemia, focal ectopies arising from this location, and including both Purkinje fibers and the surrounding working cardiomyocytes, have the highest propensity to trigger sustained arrhythmias. In conclusion, we used cell-specific optogenetics to determine with high spatial resolution and cell type specificity the requirements for the generation of extrasystoles and the factors causing ectopies to be arrhythmia triggers during myocardial ischemia.
Keywords: Purkinje fiber; arrhythmia; cardiac ectopies; heart; optogenetics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
- Channelrhodopsins for Cell-Type Specific Illumination of Cardiac Electrophysiology.Fernández MC, Kopton RA, Simon-Chica A, Madl J, Hilgendorf I, Zgierski-Johnston CM, Schneider-Warme F.Fernández MC, et al.Methods Mol Biol. 2021;2191:287-307. doi: 10.1007/978-1-0716-0830-2_17.Methods Mol Biol. 2021.PMID:32865751
- Optogenetic termination of ventricular arrhythmias in the whole heart: towards biological cardiac rhythm management.Nyns ECA, Kip A, Bart CI, Plomp JJ, Zeppenfeld K, Schalij MJ, de Vries AAF, Pijnappels DA.Nyns ECA, et al.Eur Heart J. 2017 Jul 14;38(27):2132-2136. doi: 10.1093/eurheartj/ehw574.Eur Heart J. 2017.PMID:28011703Free PMC article.
- Optogenetic defibrillation terminates ventricular arrhythmia in mouse hearts and human simulations.Bruegmann T, Boyle PM, Vogt CC, Karathanos TV, Arevalo HJ, Fleischmann BK, Trayanova NA, Sasse P.Bruegmann T, et al.J Clin Invest. 2016 Oct 3;126(10):3894-3904. doi: 10.1172/JCI88950. Epub 2016 Sep 12.J Clin Invest. 2016.PMID:27617859Free PMC article.
- Will cardiac optogenetics find the way through the obscure angles of heart physiology?Pianca N, Zaglia T, Mongillo M.Pianca N, et al.Biochem Biophys Res Commun. 2017 Jan 22;482(4):515-523. doi: 10.1016/j.bbrc.2016.11.104. Epub 2016 Nov 18.Biochem Biophys Res Commun. 2017.PMID:27871856Review.
- [Mechanisms of arrhythmias and heart fibrillation in myocardial ischemia and the possible role of Purkinje cells in their origin].Beskrovnova NN, Sharov VG.Beskrovnova NN, et al.Arkh Patol. 1992;54(6):42-5.Arkh Patol. 1992.PMID:1471932Review.Russian.
Cited by
- Novel Optics-Based Approaches for Cardiac Electrophysiology: A Review.Müllenbroich MC, Kelly A, Acker C, Bub G, Bruegmann T, Di Bona A, Entcheva E, Ferrantini C, Kohl P, Lehnart SE, Mongillo M, Parmeggiani C, Richter C, Sasse P, Zaglia T, Sacconi L, Smith GL.Müllenbroich MC, et al.Front Physiol. 2021 Nov 18;12:769586. doi: 10.3389/fphys.2021.769586. eCollection 2021.Front Physiol. 2021.PMID:34867476Free PMC article.Review.
- Optogenetics for light control of biological systems.Emiliani V, Entcheva E, Hedrich R, Hegemann P, Konrad KR, Lüscher C, Mahn M, Pan ZH, Sims RR, Vierock J, Yizhar O.Emiliani V, et al.Nat Rev Methods Primers. 2022;2:55. doi: 10.1038/s43586-022-00136-4. Epub 2022 Jul 21.Nat Rev Methods Primers. 2022.PMID:37933248Free PMC article.
- Multiphoton Imaging of Ca2+ Instability in Acute Myocardial Slices from aRyR2R2474S Murine Model of Catecholaminergic Polymorphic Ventricular Tachycardia.Borile G, Zaglia T, E Lehnart S, Mongillo M.Borile G, et al.J Clin Med. 2021 Jun 26;10(13):2821. doi: 10.3390/jcm10132821.J Clin Med. 2021.PMID:34206855Free PMC article.
- Neurohumoral Cardiac Regulation: Optogenetics Gets Into the Groove.Scalco A, Moro N, Mongillo M, Zaglia T.Scalco A, et al.Front Physiol. 2021 Aug 31;12:726895. doi: 10.3389/fphys.2021.726895. eCollection 2021.Front Physiol. 2021.PMID:34531763Free PMC article.Review.
- Optogenetic Control of Heart Rhythm: Lightly Guiding the Cardiac Pace.Dokshokova L, Pianca N, Zaglia T, Mongillo M.Dokshokova L, et al.Methods Mol Biol. 2022;2483:205-229. doi: 10.1007/978-1-0716-2245-2_13.Methods Mol Biol. 2022.PMID:35286678
References
- Sedmera D, Gourdie RG. Why do we have Purkinje fibers deep in our heart? Physiol Res. 2014;63(Suppl 1):S9–S18. - PubMed
- Hoyt RH, Cohen ML, Saffitz JE. Distribution and three-dimensional structure of intercellular junctions in canine myocardium. Circ Res. 1989;64(3):563–574. - PubMed
- Rohr S, Kucera JP, Fast VG, Kléber AG. Paradoxical improvement of impulse conduction in cardiac tissue by partial cellular uncoupling. Science. 1997;275(5301):841–844. - PubMed
Publication types
MeSH terms
Substances
Related information
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
