DNA ligase III promotes alternative nonhomologous end-joining during chromosomal translocation formation
- PMID:21655080
- PMCID: PMC3107202
- DOI: 10.1371/journal.pgen.1002080
DNA ligase III promotes alternative nonhomologous end-joining during chromosomal translocation formation
Abstract
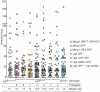
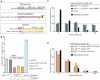
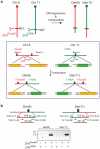
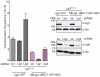
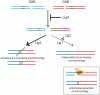
Nonhomologous end-joining (NHEJ) is the primary DNA repair pathway thought to underlie chromosomal translocations and other genomic rearrangements in somatic cells. The canonical NHEJ pathway, including DNA ligase IV (Lig4), suppresses genomic instability and chromosomal translocations, leading to the notion that a poorly defined, alternative NHEJ (alt-NHEJ) pathway generates these rearrangements. Here, we investigate the DNA ligase requirement of chromosomal translocation formation in mouse cells. Mammals have two other DNA ligases, Lig1 and Lig3, in addition to Lig4. As deletion of Lig3 results in cellular lethality due to its requirement in mitochondria, we used recently developed cell lines deficient in nuclear Lig3 but rescued for mitochondrial DNA ligase activity. Further, zinc finger endonucleases were used to generate DNA breaks at endogenous loci to induce translocations. Unlike with Lig4 deficiency, which causes an increase in translocation frequency, translocations are reduced in frequency in the absence of Lig3. Residual translocations in Lig3-deficient cells do not show a bias toward use of pre-existing microhomology at the breakpoint junctions, unlike either wild-type or Lig4-deficient cells, consistent with the notion that alt-NHEJ is impaired with Lig3 loss. By contrast, Lig1 depletion in otherwise wild-type cells does not reduce translocations or affect microhomology use. However, translocations are further reduced in Lig3-deficient cells upon Lig1 knockdown, suggesting the existence of two alt-NHEJ pathways, one that is biased toward microhomology use and requires Lig3 and a back-up pathway which does not depend on microhomology and utilizes Lig1.
Conflict of interest statement
SY-W Wong, J Lou, L Zhang, J Li, EJ Rebar, PD Gregory, and MC Holmes are full-time employees of Sangamo BioSciences.
Figures


Similar articles
- Alternative end-joining is suppressed by the canonical NHEJ component Xrcc4-ligase IV during chromosomal translocation formation.Simsek D, Jasin M.Simsek D, et al.Nat Struct Mol Biol. 2010 Apr;17(4):410-6. doi: 10.1038/nsmb.1773. Epub 2010 Mar 7.Nat Struct Mol Biol. 2010.PMID:20208544Free PMC article.
- Ligase I and ligase III mediate the DNA double-strand break ligation in alternative end-joining.Lu G, Duan J, Shu S, Wang X, Gao L, Guo J, Zhang Y.Lu G, et al.Proc Natl Acad Sci U S A. 2016 Feb 2;113(5):1256-60. doi: 10.1073/pnas.1521597113. Epub 2016 Jan 19.Proc Natl Acad Sci U S A. 2016.PMID:26787905Free PMC article.
- Marked contribution of alternative end-joining to chromosome-translocation-formation by stochastically induced DNA double-strand-breaks in G2-phase human cells.Soni A, Siemann M, Pantelias GE, Iliakis G.Soni A, et al.Mutat Res Genet Toxicol Environ Mutagen. 2015 Nov;793:2-8. doi: 10.1016/j.mrgentox.2015.07.002. Epub 2015 Jul 4.Mutat Res Genet Toxicol Environ Mutagen. 2015.PMID:26520366
- Alternative Okazaki Fragment Ligation Pathway by DNA Ligase III.Arakawa H, Iliakis G.Arakawa H, et al.Genes (Basel). 2015 Jun 23;6(2):385-98. doi: 10.3390/genes6020385.Genes (Basel). 2015.PMID:26110316Free PMC article.Review.
- Structure and function of the DNA ligases encoded by the mammalian LIG3 gene.Tomkinson AE, Sallmyr A.Tomkinson AE, et al.Gene. 2013 Dec 1;531(2):150-7. doi: 10.1016/j.gene.2013.08.061. Epub 2013 Sep 5.Gene. 2013.PMID:24013086Free PMC article.Review.
Cited by
- Exploiting DNA Ligase III addiction of multiple myeloma by flavonoid Rhamnetin.Caracciolo D, Juli G, Riillo C, Coricello A, Vasile F, Pollastri S, Rocca R, Scionti F, Polerà N, Grillone K, Arbitrio M, Staropoli N, Caparello B, Britti D, Loprete G, Costa G, Di Martino MT, Alcaro S, Tagliaferri P, Tassone P.Caracciolo D, et al.J Transl Med. 2022 Oct 22;20(1):482. doi: 10.1186/s12967-022-03705-z.J Transl Med. 2022.PMID:36273153Free PMC article.
- Removal of shelterin reveals the telomere end-protection problem.Sfeir A, de Lange T.Sfeir A, et al.Science. 2012 May 4;336(6081):593-7. doi: 10.1126/science.1218498.Science. 2012.PMID:22556254Free PMC article.
- The response to and repair of RAG-mediated DNA double-strand breaks.Helmink BA, Sleckman BP.Helmink BA, et al.Annu Rev Immunol. 2012;30:175-202. doi: 10.1146/annurev-immunol-030409-101320. Epub 2012 Jan 3.Annu Rev Immunol. 2012.PMID:22224778Free PMC article.Review.
- To indel or not to indel: Factors influencing mutagenesis during chromosomal break end joining.Cisneros-Aguirre M, Ping X, Stark JM.Cisneros-Aguirre M, et al.DNA Repair (Amst). 2022 Oct;118:103380. doi: 10.1016/j.dnarep.2022.103380. Epub 2022 Jul 30.DNA Repair (Amst). 2022.PMID:35926296Free PMC article.Review.
- Microhomologies are prevalent at Cas9-induced larger deletions.Owens DDG, Caulder A, Frontera V, Harman JR, Allan AJ, Bucakci A, Greder L, Codner GF, Hublitz P, McHugh PJ, Teboul L, de Bruijn MFTR.Owens DDG, et al.Nucleic Acids Res. 2019 Aug 22;47(14):7402-7417. doi: 10.1093/nar/gkz459.Nucleic Acids Res. 2019.PMID:31127293Free PMC article.
References
- Mani RS, Chinnaiyan AM. Triggers for genomic rearrangements: insights into genomic, cellular and environmental influences. Nat Rev Genet. 2010;11:819–829. - PubMed
- Jeggo PA. Identification of genes involved in repair of DNA double-strand breaks in mammalian cells. Radiat Res. 1998;150:S80–91. - PubMed
- Lieber MR, Ma Y, Pannicke U, Schwarz K. Mechanism and regulation of human non-homologous DNA end-joining. Nat Rev Mol Cell Biol. 2003;4:712–720. - PubMed
Publication types
MeSH terms
Substances
Related information
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
