Phylogenetic Inference
Phylogenetics is the study of the evolutionary history andrelationships among individuals, groups of organisms (e.g.,populations, species, or higher taxa), or other biological entitieswith evolutionary histories (e.g., genes, biochemicals, ordevelopmental mechanisms).Phylogenetic inference is the taskof inferring this history, and as with other problems of inference,there are interesting and difficult questions regarding how theseinferences are justified.
In this entry, we examine what phylogenetic inference is and how itworks. In the first section, we briefly introduce the field ofphylogenetics and its history. In section 2, we explore howphylogenetic inference provides useful problems for philosophers toexamine, and where philosophical approaches have contributed to thescientific examination of phylogenetics. This will help display whyphylogenetic inference is not merely a biological research problem,but a philosophical one as well. Finally in section 3, we will move toa discussion of some of the contemporary debates about foundationalissues in phylogenetics, and what the future of phylogenetic inferencelooks like.
- 1. Phylogenetic Inference in Biology
- 2. Phylogenetic Inference and Philosophy
- 3. Looking Ahead: New Challenges and Opportunities
- Bibliography
- Academic Tools
- Other Internet Resources
- Related Entries
1. Phylogenetic Inference in Biology
1.1 Primer and Introduction to Phylogenetics
A phylogeny is a reconstruction of evolutionary history. Thus thediscovery of evolution is a good starting point for the history ofphylogenetics. While Darwin was not the first to propose that somespecies were genealogically related to others, it wasOn TheOrigin of Species (Darwin 1859) that convinced many biologists toaccept common ancestry and to start building phylogenies. One of theimmediate—and ongoing—impacts was the question this raisedin how phylogenies (i.e., reconstructions of evolutionary history)relate to taxonomies (i.e., how we group taxa).
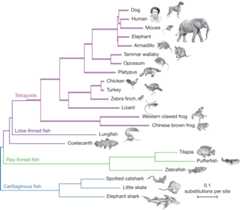
Figure 1 Phylogenetic (or evolutionary)trees have become commonplace in biology research articles. This treedisplays a recent hypothesis on the relative placement of lungfish andcoelacanth in the evolutionary history of tetrapods (Amemiya et al.2013: 312, fig. 1). [Anextended description of figure 6 is in the supplement.]
Pre-Darwinian taxonomy focused on classifying according to the“natural system” where taxa were united into large groupsdue to their “natural affinities” (Winsor 1976). In aneffort to clarify the concept of affinity, Richard Owen used the term“homologue” to refer to “the same organ in differentanimals under every variety of form and function” (Owen 1843).Put into an evolutionary framework, the natural affinities unitingthese groups were regarded as the result of descent with modificationfrom common ancestors and “homology” has since become acentral term in comparative biology (Donoghue 1992; Hall 1994). To seehow homology relates to phylogenetic inference, as well as tointroduce some basic terminology, it will be useful to consider anexample phylogeny (Figure 1).
Figure 1 provides a typical contemporary example of aphylogenetictree—a branching diagram that displays genealogicalrelationships. Phylogenetic trees like these include branches (thelines) and nodes (where the lines branch or come together, dependingon the direction you are reading the tree). Terminal branches aremarked with the entities whose evolutionary relationships are beingstudied. In this case those entities are species or higher taxa, buttrees may be constructed for other taxonomic levels, or any otherbiological entities whose evolutionary relationship can be studied(e.g., viruses, stretches of DNA, extinct taxa). Phylogenetic methodscan even be employed to study ontogeny or cultural evolution, e.g.,they have been used to construct cell fate maps to reveal how thedevelopmental parts of a single organism are related (Salipante &Horwitz 2006), and used to reconstruct the expansion of languagefamilies and help estimate historical human migration patterns (Gray& Jordan 2000; Gray & Atkinson 2003; Gray, Drummond, &Greenhill 2009). More recently, phylogenetic tools have been used toaid epidemiology studies of COVID-19 (Lemieux et al. 2021).
Since so much of modern biology requires the ability to readphylogenetic trees, good guides are commonplace. This includes journalarticles (e.g., Baum, Smith, & Donovan 2005; Baum 2008; Yang &Rannala 2012), online guides (e.g.,Understanding Evolution,Other Internet Resources), and more comprehensive textbooks (e.g., Felsenstein 2004; Lemey,Salemi, & Vandamme [eds] 2009; Wiley & Lieberman 2011; Baum& Smith 2013). Here, we provide only a brief introduction, butrecommend any of these (among others) to readers looking for a morethorough and technical introduction.
On the phylogenetic tree inFigure 1, time passes from left to right with extant groups labeled on theterminals of branches on the right.[1] Starting from the left there are two distinct branches, one leadingto Cartilaginous fish (sharks and rays), and the other to a node. Fromthat node the branches split, with one leading to Ray-finned fishes,and the other leading to another, more recent node (that ultimatelyleads to frogs, lizards, and mammals, among other taxa). Branches maybe read as representing ancestral lineages, with nodes representingcommon ancestors that diverged (or split) into distinct linealbranches.
We can also read the tree inFigure 1 by starting with the groups on the right and moving leftward, tracingbranches back to where they join at nodes. This direction takes usbackwards in time, and allows us to read off which groups are moreclosely related to others by looking at recency of common ancestry.For example, lungfish are more closely related to frogs than they areto pufferfish since lungfish and frogs share a more recent commonancestor with each other than either does with the pufferfish. To seethis, trace back along the branches from each group, starting on theright and moving left. You will converge on a node between thelungfish and frogs before reaching the node between either of thosetaxa and the pufferfish. Another way of saying this is that there is amonophyletic group (orclade) that includes lungfishand frogs that does not include pufferfish.
A monophyletic group consists of an ancestor and all of itsdescendants. Some familiar groups turn out not to be monophyletic. Forexample, if we tried to unite pufferfish and sharks into a singlegroup that excluded frogs and other tetrapods (such as the traditionalPisces) we get what is called aparaphyletic group.Individuals in paraphyletic groups all share a single common ancestor,but exclude some descendants of that ancestor. Artificial groups like“flying tetrapods” (birds plus the bats) would be calledpolyphyletic since they have multiple, distinct origins.Because paraphyletic and polyphyletic groups do not share a singlehistory that is unique just to them (i.e., there is no branch on thetree that leads just to this group), they cannot feature inexplanations in the same way that monophyletic groups can, i.e., theylack the utility and explanatory power of monophyletic groups (Velasco2008a). For example, biologists might be interested in studyingwhether flower diversity is driven by pollinator syndrome (i.e., moth,bird, or bee) or vice-versa. Phylogenetic principles predict differentpatterns of monophyly on these competing hypotheses, which can betested empirically and used by biologists as they design experimentsto test for the evidence of selection in cases like these (Whittall& Hodges 2007). This is simply unavailable without the explanatoryand organizing principles of monophyly, and one reason it has grown todominate systematics.
Each node on a tree is the origin of a monophyletic group. Forexample, the claim that the mammals form a single, united monophyleticgroup just means that all and only mammals share (i.e., are descendedfrom) a common ancestor. We have very good reasons to believe thatthis is true. For example, all and only the mammals have certaintraits (or characters) such as mammary glands, hair, and ossicles(three bones in the middle ear). Each of these traits arehomologues, i.e., their similarity is due to shared ancestry.(Homologous characters are typically contrasted withanalogouscharacters, i.e., traits whose similarity is not explained bycommon ancestry, but by some other process, e.g., convergent evolutionor reversal). The mammary glands in humans and in elephants arehomologous because the trait has been inherited from their commonancestor. Infigure 1, the ancestor of the mammals is located at the node with branchesleading to the platypus and the rest of the mammals. Once traitsemerge (as evolutionary novelties) on a branch they can be passed down(unchanged or modified) to the descendants of that lineage. Sinceplatypuses and mice have mammary glands, but no taxa stemming awayfrom earlier nodes do, we can infer that mammary glands evolved afterthe node where mammals and birds split, but prior to the mammaliannode. That is one way phylogenetic trees support inferential claims inbiology. If mammary glands had evolved prior to the node joiningmammals and birds, then we might expect to find mammary glands onbirds or lizards. Platypuses and mice are also born inside an amnioticsac, as are turkeys and lizards. However, frogs do not have anamniotic sac. This tells us that the amniotic sac evolved after thefrog lineage branched off from the other tetrapods. Parallel reasoningallows us to infer the history of the evolution of bones, tetrapodlimb structure, feathers, etc. These homologies form a nestedhierarchy of traits just as the monophyletic groups are nested, so,for example, the mammals are a part of amniotes which themselves are apart of the tetrapods. On the other hand, while the mammals and thebirds are both part of the amniotes, they do not overlap at all. Andindeed, no organisms have both mammary glands and feathers.
Explaining character distributions thus requires knowing which groupsare clades (i.e., monophyletic groups). Phylogenetic trees depictprecisely that, stating which groups are clades. This information iscalled thetopology of the tree, which conveys the relativeorder of the nodes and nothing about their absolute dating in time.Knowing which groups are clades allows us to reconstruct the evolutionof character traits, and it is the distribution of character traitsthat is the basic way that we infer which groups are clades in thefirst place. This style of reasoning (dubbed “reciprocalillumination” by Willi Hennig (1966) led to charges of circularreasoning from critics of phylogenetic inference (e.g., Sokal &Sneath 1963), whereas advocates have sought to explain why this worryis misplaced (see Wiley 1975; Sober 1988b, among many others). Infact, the nested hierarchies of traits is what led systematists toclassify taxa into a nested hierarchy in the first place, and it wasthe nested hierarchy of the taxonomic system that Darwin took to bethe most important evidence for common ancestry (Winsor 2009; Sober2009, 2010). The problems of understanding what it means for traits tobe homologous and of inferring homology will be discussed later (§2.1). Complicating things, groups of homologies do not always formunambiguous nested hierarchies, and sorting out what phylogenetichypotheses are best supported in these cases is a big part of whatgenerates debates over phylogenetic inference (more on this in§2 and§3.
Explaining character trait distributions is just the tip of theiceberg. Phylogenies are centrally important for all research inevolutionary biology. Phylogenetics lies at the heart of the linkingof the fields of systematics[2] and population genetics. Knowing a phylogeny is an important firststep to studying problems in evolutionary biology, functionalgenetics, comparative anatomy, and evolutionary developmental biology.Just as evolution is the unifying, organizing theme in biology,phylogenetics is the backbone of biological inference more generally.As Sterelny & Griffiths (1999: 379) put it, “Nothing inbiology makes sense except in the context of its place inphylogeny”.
1.2 From Darwin to Today: Three Interweaving Histories
For roughly 100 years after Darwin’sOrigin,phylogenetic research in biology was common and important. Yet,phylogenetics proceeded without much in the way of underlying theoryor explicable methodology—rather, the systematist with extensiveknowledge of some group simply relied on their judgment as to whichcharacter traits looked genuinely homologous and which charactertransformations seemed plausible. Some phylogenies were widely agreedupon due to the fact that when homologies are clear, inference iseasy. As early as the late 1800s, there was a general consensus amongsystematists around the branching order of many of the groupspresented inFigure 1. However, the relative placement of the coelacanth and lungfish wasdisputed almost immediately upon discovery of the former in 1938(Thomson 1991)—a dispute that has continued to draw attentionfrom biologists (e.g., Bockmann, De Carvalho, & De Carvalho 2013;Amemiya et al. 2013; Takezaki & Nishihara 2017).
The lungfish and the coelacanth appear key to understanding theorigins of the tetrapods, which has remained a major question in thereconstruction of evolutionary history since Darwin. But this is justone example among many. Bowler (1996) surveys the development ofphylogenetics over this time period and examines a number of thesedebates in more detail such as whether the arthropods form amonophyletic group, and the question of the origin of birds, and, inparticular, how they are related to the extinct dinosaurs.
While there are still important disagreements regarding phylogenies,the nature of these disputes have changed substantially asphylogeneticists no longer rely largely on individual expert judgment.The mid-twentieth century saw the introduction of competing accountsof the theoretical foundations of systematics (§1.2.1), a flurry of new algorithmic methods for constructing phylogenies andtaxonomic classifications, and the beginning of relatively easy accessto and employment of computational power (§1.2.3), all combined with new sources of evidence in the form of variouskinds of molecular data (§1.2.2). As biologists sought to incorporate these emerging features intophylogenetic method and theory, debates arose over how (or whether)this fundamentally changed the nature of systematics. At the sametime, debates about the foundations of systematics and the propermethods for classification and taxonomy became highly prevalent andinterwoven with arguments about these new methods and newly availabledata.
Traditionally, the emergence of phylogenetics has been presentedlargely in the context of debates between three major schools oftwentieth century taxonomic thought, often called “TheSystematics Wars” (§1.2.1). As centrally important as this was, a singular focus on that historyrisks overshadowing the impact of the molecular revolution (§1.2.2) and ever-increasing access to greater computing power (§1.2.3) on the establishment of phylogenetics. Below we consider how each ofthese twentieth century developments impacted the emergence ofphylogenetics as a distinct field of research. Yet, there are manyoverlapping themes cross-cutting these developments (often reinforcedby allegiances between individual researchers), suggesting that,ultimately, an integrated approach is needed for a rich, nuancedhistory of phylogenetics.
1.2.1 The Systematics Wars
Historically, phylogenetics emerged out of the larger field ofbiological systematics, the field of biology that studies thediversity of life and the relationships of living things through time.Today, systematists typically treat “relatedness” solelyin terms of recency of common ancestry, but this was not always thecase. Pre-Darwinian taxonomists discussed the relationships of variousgroups and their place in the “natural system”, and whilethe rise of evolutionary theory allowed that one sense of relatednesswas genealogical, it did not eliminate the idea of the broader notion.Debates about the role of phylogeny in classification and taxonomywere widespread (e.g., Huxley [ed.] 1940; Winsor 1995) though theybegan to take on a new form beginning in the late 1950s ascollaborations turned into organized research programs pushing theiragendas.
In his analysis of the period, David Hull (1988: ch. 5) titled one ofhis chapters “Systematists at War” and thus the name“The Systematics Wars” is sometimes used to describe thedebates of the period. Hull (1970) influentially compared three“contemporary systematic philosophies”, typically calledpheneticism, (ornumerical taxonomy, after theseminal Sokal & Sneath (1963) textbook of the same name),cladistics orphylogenetic systematics (afterHennig’s 1966 foundational text, an earlier version of which waspublished in German in 1950), andevolutionary systematics(following the lead of proponents like Simpson [1961] and Mayr [1969]and others).
The pheneticists disputed that we were in a position to know commonancestry with enough certainty to justify adopting it as foundationalto our notion of “relatedness”, and maintained that thetaxonomists’ job was discovering clusters of similarity (Sokal& Sneath 1963). Pheneticists were critical of what they viewed asthe entrenched approach to systematics, arguing that the traditionalapproach to selecting between hypotheses largely relied on therelative prominence or status of the biologist rather than goodscientific methodology. They explicitly framed their views in thecontext of arguments about what constituted “good science”and sought to promote independent and objective modes of assessingcompeting taxonomic groupings that relied on transparent andrepeatable methods. This included advocating for a greater separationbetween biological theory and the construction of similaritygroups.
Around the same time, another group (phylogenetic systematists)insisted that recency of common ancestry provided the besttheoretically-motivated foundation for relatedness (Hennig 1966). Likethe pheneticists, they too sought to promote good scientific methodsin systematics, and were similarly critical of established approaches.The phylogenetic systematists can be characterized by their insistencethat all taxonomic groups above the species level should bemonophyletic. Thus, if birds are descended from dinosaurs then theyare dinosaurs (Padian & Horner 2002). If the dinosaurs(or the reptiles more generally) are defined in a way that excludesbirds, then the group is paraphyletic and thus cannot be a taxon in aphylogenetic taxonomic system. Ernst Mayr derisively named them“cladists” for their obsessive focus on recovering clades(i.e., monophyletic groups) (Mayr 1965a), though its practitionerswere happy to embrace the name. When consistently carried out,phylogenetic taxonomic principles led to major revisions inlong-standing, traditional classifications—revisions that oftengenerated deep controversy (e.g., Halstead 1978; Gardiner et al.1979).
Both pheneticism and cladism were developed in opposition toentrenched views in systematics. In response, biologists committed tomore traditional methods sought to articulate the theoreticalunderpinnings of their approach, which came to be known asevolutionary systematics. With Ernst Mayr as their most prominentadvocate, evolutionary systematists sought to incorporate phylogenyinto classifications, but permitted that classifications might also(or even instead) represent important adaptive differences betweengroups (Simpson 1961; Mayr 1969). A prominent example that regularlyfeatured in these debates is the taxonomic placement of the birds andcrocodiles. Mayr argued that though birds are descended from dinosaursand reptiles more generally, birds share a cluster of adaptationswhich justify classifying them as a distinctive group at ataxonomically equivalent rank as reptiles. However, even thoughcrocodiles are more closely related to birds than to lizards and thusbelong to a clade with birds that lizards are not a part of, Mayrstill classifies crocodiles as part of the reptile“grade”—a group characterized by a well integratedadaptive complex (Mayr 1974). Thus unlike the cladists, evolutionarysystematists advocated for the inclusion of reptiles and otherparaphyletic groups in taxonomy.
One natural way to understand these debates is to recognize them assystematists sorting out “reconstructing a phylogeny” and“classifying a group of taxa” as two distinct tasks, andworking out how (if at all) these tasks ought to be reciprocallyinformative (Mayr 1965a, 1974; Griffiths 1974; Hennig 1975; de Queiroz& Gauthier 1990, 1992). With respect to this particular issue,cladistics has largely won; today when scientists designate highertaxa they are typically always describing monophyletic groups. Theprinciple of monophyly has proven to be such a powerful explanatoryand organizing tool that the commitment to it has grown widely and nowdominates systematics–in addition to informing both theory andmethodology across fields of biology This is one reason why thisperiod is sometimes referred to as “The CladisticRevolution” (Hull 1988; Kearney 2007; Haber 2009).
Yet, another framing is available. Namely, that as biologists shiftedto using phylogenies (rather than traditional taxonomies) to supporttheir explanations and justify their inferences, disagreements overtaxonomic classifications faded into the background. On this account,it is disagreements over phylogenetic hypotheses that are impactful;disputes over classifications become less consequential as they getreplaced in scientific inference by phylogenies. Joseph Felsenstein(2004: 145) dubs this the “It-Doesn’t-Matter-Very-Muchschool”, arguing that “systematists ‘voted withtheir feet’ to establish this school, long before I announcedits existence”. How phylogenetics ought to influence, inform, orconstrain taxonomic classifications is still a live debate (seeEreshefsky 2001 as well as debates over the adoption of the PhyloCode,among other debates about phylogenetic taxonomy), though one that isorthogonal to the inference of phylogenies. Contemporary debates ontaxonomic classification are largely over how to draw inferencesfrom phylogeniesfor classifications; there is verylittle dispute over the coherency of reconstructing phylogenies.Biological classification, on this account, is not viewed as acompetitor to phylogenetics, but dependent on it—and not theother way around. Phylogeneticists, like Felsenstein, can largelyproceed in reconstructing phylogenies without paying attention toclassificatory questions, though some of the metaphysical questionsabout the units of phylogenetics can come back to impact the inferenceof phylogeny itself (see§2.6.2)
As an important aside, we say ‘largely’ in the previousparagraph because—as in many cases in biology—there is notunanimity. Notably, biologists that identify aspatterncladists would dispute many of the chacterizations we haveoffered about phylogenetics, including the very first sentence of thisentry, i.e., “a phylogeny is a reconstruction of evolutionaryhistory.” Though there are a number of views that might beidentified with pattern cladism, all would typically regard that claimas unjustifiably reading a process into what is largely a claim aboutpatterns only. On pattern cladism, phylogenetic trees are typicallyregarded as graphical representations of evidence from characters; inthat regard, a phylogeny would just be a classification of thosecharacters. For the pattern cladist, directly inferring process orevolutionary history from phylogenies—or, worse, buildingevolutionary process assumptions into phylogenetic inference—isa mistake. They defend this for numerous reasons, often onphilosophical grounds, i.e., that pattern cladism embodies goodscientific inference, or that evolutionary claims require additionalinferential steps that need to be made explicit (Farris 1983). Thishas contributed to a split in the larger cladistics community. Oneupshot of this process/pattern split is that the term‘cladist’ has become ambiguous, contested, and imprecise,and it is not always clear who gets regarded as what kind of cladist(Carpenter 1987, The Editors 2016, Quinn 2017, Brower 2018a, Williamsand Ebach 2018). For example, while pattern cladists reject anyapriori assumptions about evolution (Brower 2019), so-calledprocess (or, sometimes,phylogenetic)cladists regard evolutionary claims as foundational tophylogenetics, while others (e.g.,statisticalphylogeneticists) have distanced themselves from using‘cladist’ as a label altogether (see§2.3 and the subsection§2.3.1 for more on this disputed taxonomy). We recommend that philosopherswriting in this area carefully specify the precise sense of‘cladist/m’, if they use that term at all, and cautionagainst using it interchangeably with ‘phylogenetics’.
Philosophers have largely focused on other topics in systematics thanthe pattern/process split. This is somewhat surprising as there arerich philosophical topics that could be fruitfully unpacked or drawnon as data by philosophers of science, especially for those interestedin the nature of scientific explanation and inference, or the role oftheory in biological methodology. For foundational literature onprocess versus pattern cladism see Beatty’s (1982) introductionof the designation of pattern cladism, with responses by Brady (1982),Patterson (1982), and Platnick (1982). Two good recent pattern cladisttextbooks include Williams & Ebach (2020) and Brower & Schuh(2021). There is an expansive though well contained literature withincladistics featuring debates between pattern and process cladists,e.g., Carpenter (1987), Brower (2000, 2002, 2019), and Lee (2002) arejust a very small selection of a large literature that philosophersinterested in the justification of scientific inference and methologymight explore.
Regardless of whether disputes about taxonomic philosophy died downbecause systematists came to agree or because they became lessimportant, it is a mistake for philosophers to treat the SystematicsWars as a live dispute, as opposed to an historical one. Yet, itremains an important episode in the history of systematics, and thecore conceptual debates left their mark on contemporary debates inphylogenetics (Haber 2009). At stake was not merely what notion of“relatedness” systematists ought to adopt, but whatconstituted good science in the context of inferring history andconstructing classifications, and the relative value (and evencoherence) of concepts like objectivity, testability, andrepeatability in science. But systematists no longer disputewhether phylogenetic inference may be justified as ascientific activity, but, rather, over how best to carry it out. Alarge part of why this change took place has to do with theavailability of new sources of molecular data and the introduction ofnew methods to take advantage of it. It is to this history that we nowturn.
1.2.2 The Molecular Revolution
Like Felsenstein, Sterner and Lidgard (2018) encourage historians andphilosophers of biology to “move past the systematicswars”. Their point is not that this was unimportant, but that asingular focus on it tends to overshadow the impact of other importanthistorical advances during this period. We agree. To remedy this,Sterner and Lidgard suggest paying attention to thepracticeof systematists in this period, rather than more narrowly focusing ondefinitional disputes (embodying the so-called philosophy of sciencein practice approach (Ankeny, Chang, Boumans, & Boon 2011)). Doingso reveals that alongside the Systematics Wars were the dualmethodological revolutions of the mathematization of systematics andthe incorporation of molecular data. These trends cut across theseparate schools of systematics, generating their own set ofcontroversies. This is particularly acute in phylogenetics, where therise of the use of mathematical and computational tools was bothenthusiastically embraced and deeply resisted. We start with a shortdescription of the impact of molecular data on phylogenetics, beforemoving to how mathematical and statistical tools were incorporatedinto phylogenetic methods insection 1.2.3 (for other histories on these topics see Dietrich 1994, 1998; Hagen2001; Sterner & Lidgard 2018).
Let’s first consider the appeal of including molecular data inphylogenetic analyses—perhaps even to the exclusion ofmorphological characters (e.g., Scotland, Olmstead, & Bennett 2003though see Wiens 2004). One of the most important aspects of themolecular revolution for phylogenetics was the sheer amount of newdata available to systematists, including for groups such as bacteria,where fossil data were scarce or severely lacking. Yet it was notmerely the scale and availability of molecular data that held suchappeal. As it became easier to collect, code, and collate these data,biologists argued that molecular phylogenetics provided an independentsource of evidence from morphological studies that bolstered evidencefor common ancestry and patterns of phylogeny (Zuckerkandl &Pauling 1965). In contrast, more traditional systematists such asTheodosius Dobzhanksy, Ernst Mayr and George Gaylord Simpson wereskeptical of molecular evolution studies quite generally, and, inparticular, the idea that they were in some way superior or couldreplace morphological studies (Dietrich 1998).
Many of the earliest molecular studies typically supported establishedphylogenies, e.g., Margoliash (1963) discusses the evolution ofcytochromec, though simply presumes the received phylogenyof species as correct. Later, Fitch & Margoliash (1967) constructa phylogeny solely from the cytochromec data, primarily todemonstrate molecular phylogenetic methods and how to test thereliability of results. Interestingly, one of the“anomalies” they mention is that turtles are placed closerto birds than to snakes—but this turns out to be no anomaly! Thevast majority of molecular studies have since placed the turtles as asister group to the archosaurs (a group which includes the birds andthe crocodiles) including the largest genome-scale studies done todate (Crawford et al. 2015).
The case of the phylogeny of turtles as well as other recent molecularstudies (e.g., of the lungfish and the coelacanth, see Amemiya et al.2013; Takezaki & Nishihara 2017) illustrate a general point. Evenin groups where we have extensive fossils and easily accessible livingspecimens with detailed anatomical and morphological studies,molecular evidence has often been taken to settle disputes and even tooverturn previously widely held views. This is in part due to thesheer volume of molecular data, which can overwhelm other sources ofevidence, but also because of the way molecular data may be viewed asevolving “neutrally”, i.e., as the result of mutation andgenetic drift rather than natural selection (Duret 2008), and, thus,be less prone to displayinganalogies (the result ofconvergent evolution), which can be mistaken forhomologies(the result of descent from common ancestry). (An analogy is thesimilarity relation between two analogous characters, and is a kind of“homoplasy” relation, i.e., characters whose similarity isnot due to descent from common ancestors.)
Analogies are false positives of evidence of common ancestry.Morphological traits, it is presumed, are more subject to selectivepressures that can generate these false positives, and so moleculardata from sources less subject to selective pressures (i.e., neutralsites) are preferable for reconstructing phylogenetic history. Fromthose phylogenies, analogies can be better identified, offeringevidence for appeals to natural selection (in the form of convergentevolution or other evolutionary processes) as an explanation for thepattern of biodiversity and trait distribution (Whittall & Hodges2007 provides a good example of this reasoning in regard to pollinatormode and morphology). Put another way, molecular data helps biologistsexplain why what appear to be homologies fail to form nestedhierarchies. Molecular data can identify which of those might beanalogies instead, and this helps resolve what would otherwise appearto be conflicts in phylogenetic inference. This is one way biologistsuse phylogenies to offer data-driven, theoretically-groundedevolutionary explanations, in contrast to “just-so”stories.
In addition to adding more evidence in cases where systematistsalready had a great deal of it, molecular studies can shed light oncases where other sources of evidence are extremely weak. For example,molecular data may preserve deep phylogenetic history otherwiseobscured at the morphological level by millions of years of adaptivechanges. Phylogeneticists depicting the very earliest branches inplant and animal phylogenies, as well as relationships between thedifferent groups of eukaryotes, recognized how molecular data couldboth test and enrich these deep phylogenetic hypotheses (Kenrick &Crane 1997; Baldauf 2003). Subsequent molecular phylogentic analyseshave demonstrated the fruitfulness of using molecular data (e.g.,amino acid sequences, RNA transcriptomes, and nuclear andmitochondrial genomes) for constructing deep phylogeny hypotheses(Heckman et al. 2001; Dunn, Giribet, Edgecombe, & Hejnol2014).
Emile Zuckerkandl and Linus Pauling’s introduction and use ofthe hypothesis of a molecular clock provides a good example of thepowerful utility of molecular data and was an important early episodein the history of molecular phylogenetics. The term “molecularclock” was coined in Zuckerkandl & Pauling (1965), but theyhad already used it without naming it in earlier publications(Zuckerkandl & Pauling 1962; Pauling & Zuckerkandl 1963). Thebasic idea is fairly simple. Given some estimated rate of molecularevolution, the relative ages of branches and nodes may be estimatedgiven some set of molecular data mapped onto a phylogenetic topology.Thus, molecular models of evolution were incorporated intophylogenetic methods (§1.2.3).
As more sequences of proteins became available, Zuckerkandl andPauling noted that homologous proteins displayed slight yetsystematically different sequences across taxa. For example, whilethere were 18 amino acid differences between the human and horsehemoglobin α-chains, there was thought to be only twodifferences between gorillas and humans.[3]
Zuckerkandl and Pauling then assumed that amino acid changes wereroughly linearly proportional to time, and thus the time between thedivergence of humans and horses was nine times greater than thatbetween humans and gorillas. Using paleontological estimates forcalibration, Zuckerkandl and Pauling concluded that the humans andgorillas diverged a mere 14.5 million years ago—much more recentthan traditionally was thought. This line of inference is not limitedto molecular phylogenetics, but has been imported for application toother data types, e.g., similar reasoning from phylogenetic techniqueshas been applied to language-tree divergence times to estimateIndo-European and Polynesian human migration (Gray & Jordan 2000;Gray & Atkinson 2003; Gray, Drummond, & Greenhill 2009). (Formore on the early development of the molecular clock, see Morgan1998.)
Later, it was thought that the neutral theory of evolution[4] provided some theoretical justification for the molecular clockhypothesis (Kimura 1968, 1969; Duret 2008), though Dietrich (1994)argues that the relationship goes in the other direction withZuckerkandl and Pauling’s research providing an essentialfoundation for the development of the neutral theory in the firstplace.
The molecular clock is but one of many examples of the importance ofmolecular biology to phylogenetics. Importantly, incorporatingmolecular data helped extend phylogenetic studies beyond eukaryotes,i.e., the molecular revolution has fundamentally altered (or,arguably, created) the field of microbial phylogenetics. Beginning in1969, Carl Woese began to sequence ribosomal RNA (rRNA) from acrossthe prokaryotic kingdom in an effort to understand bacterialphylogeny. The work was slow and difficult, but by 1975 Woese and hislab had sequenced the rRNA of 27 different bacterial taxa (Woese etal. 1975). In 1977, Woese and George Fox dropped a bombshell. They hadsequenced genes from a number of prokaryotic organisms whose rRNAgenes lacked the standard signatures of all previously known bacterialrRNA genes and which appeared as distantly related to the bacteria asdid the eukaryotes. Woese and Fox named them the“Archaebacteria” to emphasize their distinctness (Woese& Fox 1977). In 1990, Woese, Kandler, and Wheelis (1990) producedthe first attempt at building a universal tree of life that includedbacteria, archaebacteria, and the eukaryotes based on molecular data,with the project continuing to this day (Ciccarelli et al. 2006;Hinchliff et al. 2015; Hug et al. 2016). All of this is only possiblebecause of molecular sequence data.
1.2.3 The Mathematization of Phylogenetics and Introduction of Basic Phylogenetic Methods
The introduction of molecular data was highly significant inphylogenetics. In addition to the amount of data it generated, and theway molecular data provided evidence where other data were scarce,this new type of data was also significant in how it promoted themathematization of phylogenetics. Molecular data lends itself tomathematical and statistical analysis, in part due to (a) the ease bywhich molecular data may be encoded; (b) the systematic way molecularevolution may be mathematically modeled and subsequently incorporatedinto statistical methods; and (c) the manner by which mathematical andstatistical tools help analyze the large volume of molecular dataunder consideration. So though the molecular revolution andmathematization of systematics need not go hand-in-hand, it is hardlysurprising that they do.
A number of authors have discussed the rise of mathematical thinkingand computational tools in systematics—especially in connectionwith numerical taxonomy and phenetic classification (e.g., Hagen 2001;Sterner & Lidgard 2014, 2018). Here we focus on the rise ofstatistical methods for inferring phylogenies and later (§2) we discuss the philosophical debates that arose regarding the use ofthese methods. Good and more thorough textbook introductions forphylogenetic methods include Felsenstein (2004), Lemey et al. (2009)and Baum & Smith (2013). Yang and Rannala (2012) provide a helpfulrecent review in the context of molecular phylogenetics.)
Incorporating molecular data both encourages and is facilitated by theadoption of mathematical tools. We got a hint of that in thediscussion of molecular clocks (§1.2.2), which use estimated rates of molecular evolution to estimatedivergence times. Around the same time, Anthony Edwards and LucaCavalli-Sforza—both students of R.A. Fisher—began thinkingof phylogenetic inference as a problem for statistics. After initiallyworking independently on the problem, they collaborated to producewhat are perhaps the first set of papers presenting phylogeneticinference in a statistical framework (as attributed by (Felsenstein2001, 2004); Edwards’ own recollections of this work can befound in Edwards (1996)).
Among the earliest algorithmic methods for inferring phylogenies arethose known as “distance” methods. “Distance”could represent whichever character coding scheme we might be using toconstruct the tree, molecular or morphological, e.g., the number ofnucleotide differences between two sampled sequences of DNA. Adistance method uses these pair-wise distances to infer thephylogeny.
The first distance method developed is among the best justifiedstatistically, namely, theleast-squares method ofCavalli-Sforza and Edwards (1967). Here each topology has a set of“best-fitting” branch lengths where fit is measured bysumming the squared differences between the expected distance and theactual distance between each pair of taxa. The best tree topology isthe one whose best fit branch lengths are a better fit than any othertree’s best fit.
An alternative class of methods are known as parsimony methods. Herewe simply need a measure of how many steps it takes for one characterstate to transform into another. Recall that different tree hypothesesgenerate different explanations of the current characterdistributions. The basic idea behind parsimony is that the treehypothesis that explains these character distributions with the fewestnumber of evolutionary changes is to be preferred.
The first parsimony algorithm was published in Camin and Sokal (1965)and a number of variants of parsimony have since been developed. Forexample, one might want to weight certain kinds of changes relative toothers or even disallow certain kinds of changes (such as reversals)all together. A survey of such methods as well as a collection ofdifferent algorithms for carrying them out can be found in Felsenstein(2004). Parsimony was the chosen method for the Cladistic school whichis part of the reason that the history sketched in “TheSystematics Wars” (§1.2.1) above is intertwined with the history of mathematization sketchedhere. Philosophical arguments for and against the superiority ofparsimony as an inference method will be discussed later insection 2.3.
If we think of phylogenetic inference as an instance of statisticalinference more generally, then likelihood and Bayesian methods springto mind as methods that exemplify more general, statistical paradigms.Both require calculating probabilities and thus will require models ofevolutionary change i.e., some description of the process of evolutionand assignment of probabilities to possible changes in characterstates (such as from one nucleotide to another). Let’s brieflywalk through what that means, as how and whether to incorporate modelsof evolution in phylogenetic inference has been a major topic ofdebate.
If the data are observed nucleotide sequences from different groups ofterminal taxa, then the calculation we are after is the probabilitythat those observed sequences of nucleotides would arise on the tipsof competing phylogenetic hypotheses. The simplest way to calculatethat is to use the Jukes-Cantor model of evolution (Jukes & Cantor1969). This model assumes an equal frequency of the four nucleotides,and a uniform probability that any one nucleotide might change toanother. (These assumptions are typically violated, but reflectingthat in a model of evolution introduces additional parameters, whichincreases computing time—a valuable and limited resource in1969.) The Jukes-Cantor model provides a model of evolution on whichthe relevant probability that any given sequence might have evolvedfrom another more primitive sequence may be calculated. Different treetopologies will generate different probabilities that the observedsequences evolved from a hypothetical ancestral state. Thisconditional probability, \(P(\text{Data} | \text{Tree})\), is calledthe likelihood of the tree.
Though the Jukes-Cantor model is still used, more sophisticated modelsof evolution are available, e.g., the K80 (Kimura 1980), HKY85(Hasegawa, Kishino, & Yano 1985) and Tamura-Nei (Tamura & Nei1993) models of evolution. These introduce more parameters, e.g.,assumptions about equal frequencies of nucleotides and/or the uniformprobability of changing nucleotide states may be relaxed and evencustomized; models may also incorporate molecular clockassumptions.
These parameter-rich models reflect a better understanding ofmolecular evolution (e.g., that transversion and transition[5] rates between nucleotides are rarely equivalent) and the combinationof lowered costs and increasing accessibility to growing computationalpower that permits more parameter rich analyses. The large number ofmodels available leads to an interesting statistical problem in itsown right known as the problem of model selection, i.e., which modelof evolution do we use in our phylogenetic tools? For discussions ofthis problem in the context of phylogenetics, see (Posada &Crandall 1998; Sullivan & Joyce 2005; Posada 2008).
Once we choose a model or collection of models to work with, we cannow calculate probabilities conditional on these additional parameters(\(\theta\)). This likelihood may be formally represented as:
\[ L(\text{Tree}, \theta | \text{Data}) = P(\text{Data} | \text{Tree}, \theta) \]As a reminder, the likelihood of a hypothesis given a set of data isjust equal to the conditional probability of those data given thathypothesis. On these model-based phylogenetic approaches, the data areconditional on both a particular phylogenetic tree hypothesisand the stipulated parameters.
Using different values for the parameters in your model will lead todifferent likelihoods for the tree. The maximum likelihood estimate isthe tree hypothesis along with the fitted parameters which maximizesthis likelihood. Felsenstein (1981) gave the first computationallyfeasible algorithm for calculating likelihoods, and with moderncomputing power combined with the huge amounts of sequence dataavailable, likelihood methods have become the most commonly usedmethods in phylogenetics.
Just as with likelihood methods, Bayesian methods require a model ofevolution before any inferences are possible. The fundamentaldifference is that likelihood inference treats parameters of the modelas nuisance parameters which have a fixed but unknown value. In thefrequentist method of maximum likelihood, these parameters are set totheir best fitting value. Bayesian inference treats parameters asrandom variables with unknown distributions and probabilities are usedas a measure of uncertainty. The probability of a tree is aconditional probability \(P(\text{Tree} | \text{Data})\) under theassumption of a particular model. When we incorporate the modelparameters \(\theta\), by Bayes’ theorem, we have:
\[P(\text{Tree}, \theta | \text{Data}) = \frac{P(\text{Data} | \text{Tree}, \theta)*P(\text{Tree}) }{P(\text{Data})}.\]Controversially, this requires that we attach prior probabilities tothe possible hypotheses as well as the parameters in our model.Pickett and Randle (2005) claim that it is impossible to assign priorsin a consistent way. Velasco (2007) argues that this claim rests on amistake, though the problem remains a difficult one. He lateradvocates assigning priors to tree topologies, but leaves open themore difficult problem of priors on branch lengths or model parameters(Velasco 2008b). Alfaro and Holder (2006) attempt to address some ofthese issues.
Anthony Edwards (1970) was the first to discuss Bayesian methods inphylogenetics, but it was computationally impossible to carry theseinferences out at the time. In the 1990s, three groups independentlydeveloped methods for carrying out Bayesian inferences of phylogeniesin practice (Rannala & Yang 1996; Mau & Newton 1997; Yang& Rannala 1997; Li, Pearl, & Doss 2000). Each group usedBayesian Markov chain Monte Carlo (MCMC) algorithms to estimateposterior probabilities, which other groups further refined (e.g.,Larget & Simon 1999; Ané, Larget, Baum, Smith, & Rokas2007; Huelsenbeck, Ané, Larget, & Ronquist 2008). (SeeArchibald, Mort, & Crawford 2003 for an early and non-technicalprimer on Bayesian phylogenetic methods.) With further statistical andcomputational improvements, Bayesian methods are now fairlycommonplace in the literature alongside likelihood methods.
2. Phylogenetic Inference and Philosophy
Section 1 served as an introduction to the history and problem of phylogeneticinference. In this section, we will look at how phylogenetics andphilosophy are intertwined—in particular, we will examine somefoundational debates in phylogenetics that have a distinctlyphilosophical flavor, and we will point out more traditionalphilosophical questions that the study of phylogenetics can shed lighton.
At its core, phylogenetic inference is about evaluating competinghypotheses. In an important sense, phylogeneticists are faced withwhat philosophers of science would identify as a problem ofunderdetermination of theory by evidence .Multiple competing phylogenetic trees can explain the same data,though in conflicting ways; it is the phylogeneticists task toidentify which of those hypotheses best explains those data. It willbe useful to separate the problem into two parts:
- identifying what the evidence is; and
- constructing a phylogeny to explain the evidence we have.
In the case of phylogenetic inference, we are in the philosophicallyinteresting and puzzling situation where it seems that these twoquestions cannot really be separated.
2.1 Alignment and Character Coding
The question of which method of phylogenetic inference is bestjustified is clearly important, but skips over an important step inphylogenetic inference—the construction of a data matrix ofcharacters. Mishler (2005), for one, argues that it is the mostimportant step! Following Winther (2009), we can separatecharacter analysis which results in the construction of adata matrix fromphylogenetic analysis which treats the dataas input and produces a phylogeny. This first stage of characteranalysis has been widely discussed in biology (though surely lesswidely discussed than the second stage). In addition to the voluminousliterature on homology, useful collections of work on charactersinclude those edited by Wagner (2000) and Scotland & Pennington(2000).
Character analysis includes identifying characters of the organisms weare looking at, and in particular, identifying when a character in oneorganism is the same character as in another organism. This is theproblem of homology. A different, but related problem, is assigningcharacter states. For example, one bear might have black fur whileanother has white fur. Here, “fur” is the character, butit is in a different state (fur color) in the two organisms. Though itis standard to treat characters and character states separately, theextent to which these are genuinely different problems rather thanthan aspects of the same problem is a matter of debate (Stevens 2000;Freudenstein 2005; Sereno 2005).
Identifying and coding characters is certainly a biological problem,but it is a philosophical one as well. On what basis can we judge thatone character is homologous to another? Debates on this extend back tothe very beginning of contemporary biology, e.g., Richard Owen’sinfluential pre-Darwinian notion of homologue to Ernst Haeckel workingto develop accounts of homology in the context of Darwinian evolution.In the mid-twentieth century, the pheneticists argued that characteridentification must be “theory-free” and advocated usingraw similarity as a guide to identifying homology (Sokal & Sneath1963; Sneath & Sokal 1973). There are well known philosophical andbiological problems with trying to use similarity in this way (e.g.,Goodman 1972), many of which were raised as objections to strictpheneticism during the Systematics Wars (e.g., Mayr 1965b; Neff 1986;Rieppel & Kearney 2002 though see Lewens 2012 for a recent defenseof pheneticism in a contemporary context).
In a phylogenetic context, homologies are regarded as the result ofdescent with modification from common ancestors. Similarity andidentity, in that context, is an expression of shared evolutionaryhistory. At the same time, homologies are also treated as hypothesesfrom which we can infer phylogenetic hypotheses (and, subsequently,both test and explain our hypotheses of homology). Clearly, even in acontemporary phylogenetic context, questions remain on how toindividuate characters and identify homologies. Richards (2002, 2003)worries that since there is no algorithm for individuating characters,we rely on illegitimate, nonscientific factors rendering characteridentification and phylogenetic analysis ultimately subjective.Rieppel and Kearney (2007) and Winther (2009) respond that characteranalysis can be considered objective since it relies on various kindsof causal criteria; Kendig (2016) adopts a slightly differentstrategy, rejecting the idea that we need to develop an analyticdefinition of “homology”, instead looking to biologicalpractice as a guide for individuating characters.
While character analysis and phylogenetic analysis are logicallyseparate tasks, in theory they should be intertwined. A givenphylogeny can make an assessment of homology more or less plausible.One proposal to avoid this problem is the introduction of adistinction between “primary” homologies (an initialcharacter identification based on structural features) and“secondary” homologies (similarity due to common ancestry)which are inferred through phylogenetic analysis (Pinna 1991; seeRoffe 2020 for a recent philosophical treatment of this distinction).On the other hand, Rieppel and Kearney (2002) argue that characterhypotheses need to be independently testable and phylogenetic analysiscan clearly provide evidence for miscoding of characters—even ifthis is not the intended goal.
Molecular data presents a different set of challenges forindividuating characters and coding them into a matrix, nicelyillustrated by DNA sequence data. With only four base nucleotides,coding is easy (§1.2.3), and modern laboratory techniques provide highly reliable sequences ofnucleotides in a stretch of DNA. But here, the problem isalignment. How do we know that one stretch of nucleotidesequences should correspond to similar stretches in other taxa? Wemight consider chromosomal location, or even functional similarities,though each of these raise other questions of how we decide what oughtto be treated as homologous characters (even contingently) acrosstaxa. This becomes even more complicated as we consider how historicalprocesses like gene duplication/loss or recombination add complexityto these questions. This is the problem of positional homology. Likemorphological character coding, molecular alignment is intertwinedwith phylogenetic analysis and various methods of joint inference havebeen proposed (Redelings & Suchard 2005; Wheeler et al. 2006).
2.2 Computational Limits and Big Data
Bearing in mind what we said above about the interconnectedness ofcharacter analysis and phylogenetic analysis, we will now proceed tothe second part, analysis of the data. At this stage, computationallimits loom large. Even if we are concerned only with inferring thetopology of a phylogeny, the number of possible trees growsexponentially relative to the number of terminal branches we includeon a tree. The formula for calculating the number of possible treetopologies is relatively straight-forward (Felsenstein 2004):
\[\frac{(2n-3)!}{2^{n-2}(n-2)!}\]wheren is the number of taxa. The upshot is that even asmall expansion of terminal branches results in an exponentialincrease in the number of possible topologies. So, for example, thereare 15 possible phylogenetic trees for a group of 4 species, over 34million for 10 species, and \(8 \times 10^{21}\) with 20 species.[6]
This means that exhaustive evaluation of all possible hypotheses veryquickly becomes all but impossible—even as our capacity for thishas grown with increased computing power. Finding the mostparsimonious tree, the maximum likelihood tree, and even doing amultiple sequence alignment to begin with are all NP-complete problems[7] (Graham & Foulds 1982; Wang & Jiang 1994; Chor & Tuller2005). Bayesian inference is similarly computationally intractable(Ronquist, Mark, & Huelsenbeck 2009).
In response, researchers have developed heuristic search strategies toexplore possible tree space. For example, in Bayesian analysis theprobability space of possible trees is so large that starting byrandomly selecting a tree will be highly inefficient. It will almostsurely fail to provide a good estimate of the posterior probabilitydistribution, since the interesting regions will only occupy a smallpart of that distribution space (Ronquist et al. 2009). A commonheuristic is to start with a parsimony or distance-methods tree.Though these methods typically perform worse than model-basedapproaches, they are much less computationally expensive and canquickly generate an initially plausible tree that Bayesianphylogeneticists use as a launching off point to explore the treeprobability space (e.g., using hill-climbing algorithms such as MarkovChain Monte Carlo (MCMC) sampling). Innovations like these are onereason that computational phylogenetics is an important field in bothstatistics and computer science. Philosophers of science interested inhow scientists explore and use so-called “Big Data” wouldprofit from examining how phylogeneticists have developed, adapted,and employed heuristic search strategies.
Setting aside computational and practical problems leaves us with whatappears to be a straightforward epistemological question—whichmethods are best justified in licensing phylogenetic inferences? Webegin our examination of this question by looking at arguments infavor of a particular method, parsimony.
2.3 Justifications of Parsimony
Parsimony methods were among the very first adopted by cladists. Thisis often taken to include Hennig (1966), who laid out a process ofphylogenetic inference that is now referred to as “Hennigianreasoning” in his seminal textPhylogenetic Systematics(Baum & Smith 2013). Like other forms of parsimony, this methodsought to identify the tree that explained the distribution ofhomologies by appeal to the fewest number of evolutionary events.
Parsimony techniques have grown more sophisticated, and, as largerdata sets have been assessed and bigger trees constructed,justifications of parsimony approaches advanced and adapted. Early on,much of this took place in the context of the Systematics Wars (§1.2.1), where a premium was placed on defending approaches as properlyscientific. Thus, these debates tended to veer into philosophy ofscience, with systematists disputing what counted as doing goodscience in the context of inferring history. This helps explain theinfluence ofKarl Popper’s work. Earlyproponents of parsimony were attracted to Popper’sfalsificationist account of the scientific method, both in the way itprovided demarcation criteria for scientific activity, and in the wayit was positioned as critical oflogical positivist approaches. Falsificationist justifications were employed to criticizenumerical taxonomists as verificationists relying on flawedhypothetico-deductive methods.
Wiley (1975) provides a representative example of a falsificationistdefense of parsimony that proved deeply influential, e.g., Eldredge& Cracraft (1980) adopt Wiley’s core justification in theirinfluential textbook, as does Farris (1983) in his impactful defenseof parsimony. On Wiley’s account, each taxonomic character istreated as an homology, and considered independently of all others.For each homology, a tree hypothesis is generated that requires thefewest number of evolutionary changes to explain the pattern ofcharacter distribution across taxa. These are hypotheses of homologyand, on the falsificationist account, these hypotheses are mutuallytestable. That is, the tree hypotheses generated for each characterconstitutes a test of the trees hypothesized for every othercharacter; falsified hypotheses of homology are treated as ad hocassumptions of homoplasy. The maximum parsimony tree is that topologywhich generates the fewest falsified hypotheses of homology, and isregarded as the most corroborated tree. Adding new characters or taxaconstitutes further testing of these hypotheses, and generates morepotential falsifiers. In this way, parsimony is simultaneously a modeof tree construction and evaluation (Haber 2009); Wiley (1975),Eldredge and Cracraft (1980), and Farris (1983) describe this as alogical basis of reasoning about phylogenetic inference, citing thephilosophical justification of falsificationism.
Whether this represents a faithful account of Popper’sfalsificationism, or whether falsificationism representstheoptimal (or even a desirable) mode of scientific reasoning has been amatter taken up by philosophers (Hull 1983, 1999; Sober 1988b), thoughthat is, in some ways, beside the point. (For a philosophically-mindedbiologist’s survey of the role of Popper in systematics, seeGaffney 1979, Rieppel 2008, or Santis 2020.) Regardless,characterizing parsimony in falsificationist terms provided animportant justification that resonates to this day. More recentinvocations of Popper in phylogenetic inference have focused onwhether parsimony or model-based methods such as Maximum Likelihood (§1.2.3) better align with falsificationism (e.g., Siddall & Kluge 1997;de Queiroz & Poe 2001, 2003; Kluge 2001), and whetherprobabilistic and statistical reasoning is relevant for studyingsingular historical events (Siddall & Kluge 1997; Haber 2005).(See Turner 2007; Cleland 2011;Currie 2018, 2019, for generalphilosophy of historical science.) Insection 2.4 we describe how Felsenstein’s (1978) discovery that parsimonyis prone to systematic errors poses a serious challenge to thisunderlying falsificationist justification.
Parsimony can be defended without falsificationism, and otherjustifications have been offered that appeal to the method’snamesake, i.e., the principle of parsimonious reasoning or appeals toOckham’s Razor (e.g., Kluge 2005). It amounts to (i) defendingparsimony (the principle) as the criterion by which we ought to assesscompeting phylogenetic hypotheses, and (ii) arguing that parsimony(the method) is the best mode for achieving that goal by seeking outthat tree that minimizes the number of evolutionary novelties requiredto explain the distribution of characters. This may also be presentedas identifying that tree that minimizes the number of ad hocassumptions of homoplasy. Though this appeal to simplicity may bewedded to falsificationist justifications (as we saw above), they arelogically independent defenses. There is certainly an intuitive appealto the principle of parsimony in the sciences, and Ockham’srazor remains influential for good reason. But justifying parsimonymethods by appeal to simplicity is not so straight-forward. Evaluatingwhich phylogenetic approaches are justified by Ockham’s Razordepends on what is being counted (Haber 2009). Sober (1988b, 2015among others) provide useful discussions of the role of simplicity andthe principle of parsimony in phylogenetics.
Other philosophical justifications of parsimony have been offered.Fitzhugh (2008 and elsewhere) has argued that the problem ofphylogenetic inference is best understood as an abductive problem, andthat treating it as a problem for statistical (or inductive) inferenceamounts to a category error. Sober (1988b) agrees that phylogeneticsinvolves abductive reasoning, though argues that thisjustifies the use of statistical approaches (especiallylikelihood).
It is also worth recalling why pattern cladists in particular objectto model-based methods of phylogeny reconstruction. Pattern cladiststypically rejectany method of phylogeny reconstruction thatincludes assumptions about the evolutionary process (§1.2.1),e.g., Brower (2019: 717) identifies a centrral commitmentdistinguishing pattern from process (or traditional or phylogenetic)cladism as the rejection of “a priori evolutionarybackground assumptions from the inference of patterns of relationshipsamong taxa.” As such, pattern cladists have strongly objected tolikelihood and other model-based phylogenetic methods, as thoseexplicitly incorporate evolutionary models (§1.2.3 and§2.4). This has typically gone hand-in-hand with a rejection ofstatistical approaches more generally, and a commitment to parsimony,e.g., “the statistical approach to phylogenetic inference waswrong from the start, for it rests on the idea that to study phylogenyat all, one must first know in great detail how evolution hasproceeded. That cannot very well be the way in which scientificknowledge is obtained” (Farris 1983: 17).
In response, statistical phylogeneticists have argued that parsimonymethods implicitly include evolutionary models, which model-basedmethods simply make explicit and, thus, subject to interrogation (deQueiroz & Poe 2001, 2003; Swofford & Sullivan 2009). Thisfault line is further reflected in the way pattern cladists view otherproponents of parsimony as allies in this debate, as Brower (2019:717), approvingly citing Kluge & Farris (1999), makes clear:“despite this difference of opinion regarding criticalbackground knowledge, there is no practical distinction betweenpattern and process cladists’ evidence, analyses or results.From a methodological perspective, all cladists group by synapomorphyalone.” That is, it is in the interpretation of methodology andinferential justification that distinctions are drawn between patternand process cladists; in the larger debate with statisticalphylogeneticists they are allied. Note, however, that ever finerdistinctions of the term ‘cladist’ are being introduced(let alone ‘statistical phylogeneticists’, who typicallyassociate with model-based approaches). This reflects the way thisterm has grown contested and associated with various competing sets ofcommitments. It is worth taking a brief detour in section 2.3.1 tosort through this a bit.
2.3.1 Who Owns the Term “Cladist”?
Insection 1.2.1, we used the term “cladistics” to refer to a school oftaxonomy which held that higher taxa must be monophyletic, and stoodin contrast to the competing schools of phenetics/numerical taxonomyand evolutionary taxonomy. In this historical context,“cladistics” was sometimes used interchangeably with“phylogenetics”. We caution against that practice,especially when discussing more contemporary practices. Though it ishard to pin down a precise date when these terms began to carrydifferent connotations, Felsenstein (1978) and Beatty (1982) are asgood a marker as any for when these terms began to diverge and take onmore nuanced senses. This is particularly true of the term“cladist”, which has become a highly disputed term, andone that has been claimed by sub-groups within the larger field ofphylogenetics.
An apt example of this is the journalCladistics, whoseeditors published a controversial editorial in 2016 stating thatsubmitted manuscripts should reflect the philosophical commitments ofthe journal and its sponsoring professional society (The WilliHennig Society) (Editors 2016). They write,
The epistemological paradigm of this journal is parsimony. There arestrong philosophical arguments in support of parsimony versus othermethods of phylogenetic inference (e.g., Farris 1983).
The editorial continues, stipulating that “Phylogenetic datasets submitted to this journal should be analysed usingparsimony”. If trees generated by alternative methods showdifferent results, these may be included if authors prefer thosetopologies, but only if authors are “prepared to defend it onphilosophical grounds” (all quotes from Editors 2016: 1). Theeditors, here, are using the imprimatur of the journalCladistics to stake out a specific sense of how“cladistics” ought to be understood, and what callingoneself a “Cladist” signals about the underlyingtheoretical and philosophical commitments held about phylogeneticinference, i.e., an exclusive commitment to parsimonymethods—going so far as to reject model-based statisticalapproaches entirely.
Aleta Quinn (2017) argues that the term “cladist” remainsone claimed by multiple overlapping (yet importantly distinct) camps.She tackles the ambiguity head on, providing a useful “foreignlanguage dictionary” of ways that “cladist” getsused in systematics. Quinn argues that the term is in need ofdisambiguation, drawing seven different senses from the literature.This includes systematists that advocate adherence to particularphylogenetic methods (e.g., parsimony, as in the case of theCladistics editorial) or philosophical justifications(especially to a version of Popperian falsificationism), though thevarious distinctions of competing senses of “cladist” gowell beyond this. In response to Quinn, Williams and Ebach (2018)propose an eighth (and their preferred) sense of cladist, and Brower(2018a) offers a focused commentary on Quinn’s article, as wellas other attempts to clarify the philosophical foundations andcommitments of pattern cladism. For example, a distinguishing featureof pattern cladism is the rejection ofa priori assumptionsabout evolutionary theory in phylogenetic methods. This has beeninterpreted as a claim of theory-neutrality or minimality by somephilosophers and biologists (e.g., Pearson 2010). Brower (2019)contests this interpretation, arguing that the claim of theoryneutrality is specific to evolutionary assumptions, as opposed to amore general claim.
The primary take away is that care should be taken when philosophersuse the term “cladist”. Though there are a lot ofinteresting philosophical topics to unpack around this term, it mustbe acknowledged that it has become ambiguous. There are simply toomany senses of this term for it to be used without clearly specifyingprecisely what sense is being invoked. We encourage philosopherswriting about phylogenetics to follow the lead of systematists, andavoid using the terms “cladist”, “cladistics”,and other cognates as synonymous with “phylogenetics”.Instead, philosophers should reserve usage of those terms andcarefully specify which sub-set of phylogeneticists (or their work)those terms are referring to.
2.4 Long-Branch Attraction and model based methods
Insection 2.3 we mentioned that the falsificationist justification of parsimony wasmet with a serious challenge by the discovery that it was subject tomaking systematic errors under certain conditions. JosephFelsenstein’s (1978) paper, “Cases in which Parsimony orCompatibility Methods Will be Positively Misleading” firstdescribes what came to be known as the phenomenon of“long-branch attraction”. Though hardly the beginning ofdebates between parsimony and model-based phylogenetic methods, itrepresents an important inflection point in discussions aboutphylogenetic inference.
Long-branch attraction refers to the tendency of some methods topreferentially group together long branches (those with moreevolutionary changes) regardless of their actual history. Methodsdesigned to identify trees requiring the fewest number of evolutionarychanges to explain the data will prefer explanations of sharedancestry over convergent evolution—even in cases whereconvergent evolution occurred. For example, imagine two branches withrapidly evolving DNA sequences. By chance alone, the two branchesmight have the same mutation at the same site which parsimony treatsas evidence that these branches share a recent history. In a range ofcases, the chances of these similar mutations occurring is so highthat we expect the number of these convergent cases to overwhelm thesignal from the true homologies. In these cases, the parsimony methodis statistically inconsistent. A method is statistically consistentwhen it is guaranteed, as more data are added, to identify the correctsolution. Methods that fail to do this are statistically inconsistent.In this case, not only does parsimony fail to be consistent, but themethod reliably returns increased support for a specific, incorrectoutcome as more data are added and hence is “positivelymisleading”.
Long-branch attraction is more than just an operational problem, italso challenges the falsificationist underpinnings of parsimony (Haber2009). Recall that on Wiley’s (1975) influential account,phylogenetic trees are subject to being falsified as more data(characters) are discovered or added (§2.3). If a more parsimonious tree is available to explain those data, thepreviously most parsimonious tree is treated as falsified (or ascontaining more ad hoc hypotheses of homoplasy). Felsenstein’sdiscovery was that in some circumstances, as more characters areincluded in a parsimony analysis, parsimony will be prone to rejectinghypotheses that report the actual phylogenetic relations, whilecorroborating more parsimonious—but incorrect—phylogenetichypotheses. This undermines the logical justification of parsimonyanalysis.
Felsenstein (1978) was (and remains) an immensely impactful paper,generating numerous research programs. It is a largely theoretical andabstract paper that considers how different methods perform underdifferent biological conditions. One natural question that might beposed concerns how prevalent these conditions are in the actualsystems being studied. This is an empirical question, of course, andcan be studied as such, though disputes over how to interpretempirical cases have proven controversial (Huelsenbeck 1997; Whiting1998; Siddall & Whiting 1999; Yang & Rannala 2012).
In the 1990s experimental phylogeneticists used empirical andsimulation studies to more rigorously empirically test claims ofperformance of competing phylogenetic methods. This included testingcompeting methods to reconstruct known in-vivo phylogenies (ofbacteriophage) that were carefully constructed, tracked, and archivedover many generations and more complex simulated phylogenies (takingadvantage of the ever-growing processing power available on the labbench) (Hillis, Bull, White, Badgett, & Molineux 1992; Hillis& Bull 1993; Hillis, Huelsenbeck, & Cunningham 1994). Theselargely confirmed that distance and parsimony methods were subject tolong-branch attraction, where model-based statistical methods such aslikelihood avoided these epistemic traps.
2.5 Phylogenetics & Philosophy of Statistics
Felsenstein’s (1978) paper is also impactful because of hiscommitment to statistical consistency asthe standard bearerfor evaluating competing statistical approaches. For Felsenstein andmany other phylogeneticists, showing that parsimony is statisticallyinconsistent is a fatal blow. But not everyone agrees. In fact, someauthors argue that it is close to an irrelevant consideration. We nowturn to this debate about the importance of consistency as an exampleof how philosophy plays a central role in phylogenetic theory.
Like Edwards and Cavalli-Sforza before him, Felsenstein (1978) treatsphylogenetic inference as a problem for statistics. Jerzy Neyman(1971) pointed to the emerging field of molecular phylogenetics as asource of novel and interesting statistical problems. In theintervening fifty years the field has only expanded in importance tothe point where phylogenetic inference is now one of the centralproblems in statistics. Similarly, research on algorithms forinferring phylogenies has been important in computer science fordecades.
If phylogenetic inference is seen as a problem in statisticalinference, it might be thought that general arguments for idealstatistical methods will just apply in this case. Maximum likelihoodor Bayesian methods could be defended on these grounds for example. Inthis light, parsimony is a statistical method and as such, we canstudy its statistical properties (Felsenstein 1983, 2004). AnthonyEdwards (1996) claims that his initial usage of the “minimumevolution” principle was as an approximation to the maximumlikelihood solution which he believed to be justified on general,statistical grounds, i.e., parsimony methods could be assessed as astatistical approach. Since that time, a number of authors have shownexact connections (ordinally equivalent rankings) between parsimonyand likelihood in a range of cases (Felsenstein 1973; Sober 1985;Tuffley & Steel 1997; Steel & Penny 2000).
In an exchange on parsimony and likelihood between Felsenstein andElliott Sober, Felsenstein defends the idea that consistency “isa fundamental property” and in particular, that it is morefundamental than likelihood (Felsenstein & Sober 1986). Hereiterates his earlier claim that
maximum likelihood methods are not desirable in themselves, butbecause they have desirable statistical properties such as consistencyand asymptotic efficiency. (Felsenstein 1978: 408)
With this attitude, Felsenstein is merely following an old andvenerable position in statistics. For example, Fisher (1956: 141)called consistency “the fundamental criterion ofestimation” and said that inconsistent estimators are“outside the pale of decent usage” (Fisher 1935 [1950:11]). Neyman (1952: 188) agrees.
Contrary to Felsenstein’s position, Sober claims that
One does not “justify” a method by showing that there isan extremely special case in which it does its work well; nor does one“refute” a method by showing that there is another specialcase in which it makes a hash of things. (Sober 1988b: 76)
One particular argument that he gives against consistency is that itcan conflict with likelihood. Sober (1988a) presents an example inwhich likelihood inference can fail to be consistent. But Soberfollows Anthony Edwards (1972) in positing that likelihood is a“primitive postulate” which does not need justificationbased on repeated sampling. If a likelihood inference can fail to bestatistically consistent but is still justified, then clearlyconsistency cannot be a necessary criterion of justifiedinference.
A different kind of worry about consistency is expressed by authorswho point out that it is not clear what consistent method we coulduse. As the editors ofCladistics put it in their editorialdefending the use of parsimony,
we do not consider the hypothetical problem of statisticalinconsistency to constitute a philosophical argument for the rejectionof parsimony. All phylogenetic methods, including parsimony, mayproduce inconsistent or otherwise inaccurate results for a given dataset. (Editors 2016)
Maximum likelihood estimation is known to be provably consistent undera wide variety of conditions (Wald 1949), but several authors haveargued that these conditions do not apply to estimating the treetopology since tree topologies are discrete, not continuous,parameters (Yang 1996; Siddall 1998; Farris 1999). However, Swoffordet al. (2001) argue that Wald’s conditions do apply and Yang(1994), Chang (1996), and Rogers (2001) prove that maximum likelihoodis consistent under different assumptions about character evolution.But regardless of which assumptions suffice for likelihood to beconsistent, some assumptions about the evolutionary process aredefinitely required. But what happens when the model we use is not thecorrect model? This is especially problematic when combined with theview that to be a correct model it should be true, and that we couldnever know that our model was true or perhaps even that there is nosuch thing as a true model, as some pattern cladists have claimed(e.g., Brower 2016, 2018b). As Felsenstein (2004: 272) puts it,
likelihood is usually consistent if we use the correct model in ouranalysis. When we use the wrong model, there are few guarantees.
2.6 Phylogenetics and Philosophy
The philosophy of statistics is but one area among many wherephilosophical arguments contribute to work in phylogenetics. Thoughphylogenetics is a subfield of biology (and, arguably, of statisticsand computer science), phylogenetic inference is a topic on which bothphilosophers and biologists work, often in overlapping ways. Indeed,the boundary between science and philosophy here is blurry in a wayreminiscent of some of the natural philosophy of the seventeenth andeighteenth centuries. An upshot of this is that phylogenetic inferenceis an enormously rich area for philosophers to explore. Biologists areactively debating the conceptual foundations of phylogeneticmethodology, justification, goals, explanatory models, etc. It is notuncommon to see philosophers playing a central role in these debates(e.g., Hull 1988; Sober 1988b). It also means that many biologists areeager to engage with philosophers in meaningful and deeply informedways (e.g., Sterner & Lidgard 2018; Havstad & Smith 2019).
Phylogenetic inference is, generally, best categorized as a problem ofepistemology. It is, after all, a problem of inference, albeitspecialized to how we justify or ground our beliefs about phylogeny.An epistemologist who wonders how we can have knowledge of the pastcan hardly do better than to look to phylogenetics (Sober 1988b). Andas we have seen, philosophy of statistics and of inductive inferencemore generally looms large. But moving beyond epistemology, anyphilosopher interested in how science works can learn fromphylogenetics. Here we will look at the cases of metaphysics and thegeneral philosophy of science.
2.6.1 Phylogenetic inference as general philosophy of science
Understanding details of the evolutionary history of life is notsimply a stand-alone problem (as interesting as that problem wouldbe). The history of life and the history of the planet are tiedtogether such that knowledge in one domain aids our inferences in theother. Knowing what types of fossils are embedded in rocks togetherwith a phylogeny can help scientists date the rocks just as dating therocks through independent means aids in dating the fossils and thus inestimating aspects of the phylogeny (Grantham 2004). And just as thephysics and chemistry of the planet have shaped living things, lifehas shaped the planet as well. Photosynthetic cyanobacteria werepartially responsible for the great oxygenation event roughly 2.4billion years ago, where oxygen build up in the Earth’satmosphere fundamentally altered the chemistry and physics of theearth.
This entanglement with other sciences makes phylogenetics a richsource for philosophers of science. It provides a model to betterunderstand how science works when specialized fields inform and relyon each other. Philosophers might draw on phylogenetics to betterunderstand the social structure of complexes of sciences (Longino1990, 2019) (and complex sciences that draw on other fields), or tocharacterize what constitutes good scientific activity for what mightbe called “service sciences”, i.e., sciences whoseproducts are largely used by other scientists as end-users.
One way to frame debates in the Systematics Wars is as scientistswrestling over the question of what constitutes good science. Here wetake the opportunity to highlight what may be an under-appreciatedaspect of this debate. Namely, does good science require theory?Though many of the traditional sciences studied by philosophers ofscience are clearly driven by (or, at least, tightly linked to)theory, this does not always appear to be the case for other sciences.Contrast Newtonian mechanics or population genetics with, say, thestudy of stem cells (Fagan 2013) or developmental biology (Love 2014),where it is unclear whether there areany underlyingtheoretical structures.
In the case of systematics, it is unclear how to articulate aprecise theory for any of these competing views. It isprobably more accurately portrayed as competing sets of commitments,some of which may resemble (or be) scientific theories (depending onthe characterizations of “scientific theory”), where othercommitments might be better described in terms of scientific practiceor mode of reasoning (Giere 1979 [1997]; Griesemer 2000).
Whether and what the theoretical structure of phylogenetics is remainsan open debate. Yet, though scientists that research, study, anddevelop phylogenetic methods have a lot at stake in these debates,most of the endusers of these methods are rather agnosticabout those competing methods (and, in many cases, the underlyingtheories supporting those methods). In practice, most biologists thatuse phylogenetic methods typically use lots of them.Philosophers of science might consider whether this amounts to anadoption of methodological pluralism or theory agnosticism, andwhether trees produced under competing methodologies constitute robustresults (e.g., when identifying the same monophyletic groups orhomologies).
Phylogenetic inference should also be of interest to a wider range ofphilosophers for the way it can function as a case study for otherphilosophical issues. The recent history of phylogenetics has providedphilosophers with test cases for how scientists shift from one set ofcommitments to another (Hull 1988), for the role of inductive logicand probability in both explanation and model construction (Sober1988a,b; Haber 2005), and for what constitutes a good scientificexplanation (Felsenstein & Sober 1986; Sober 2004; Haber 2009).O’Hara (1988) and Ereshefsky (2012) look to phylogenetics to seehow narrative or historical explanations function in the sciences.Other examples of philosophers using phylogenetics to assessphilosophical theses include Velasco (2012) and Vassend (2020) usingphylogenetics to examine practices of modeling and idealization, andHaber (2005) looking at what phylogenetic inference tells us of howprobabilistic reasoning may be used to assess singular historicalevents. Aleta Quinn proposes a taxonomy of disputes in contemporarysystematics that she argues stems from disputes over application ofthe principle of total evidence (2019), and uses the case ofphylogenetics to examine a well known objection leveled againstproponents of inference to the best explanation in the sciences(2016), the “best of the bad lot argument” (van Fraassen1989), or the “underconsideration objection” (Lipton2004). And finally, Andreasen (1998, 2000) looks to phylogenetics toanswer the question of whether biological races are real.
2.6.2 Phylogenetic inference as a problem of metaphysics
Though phylogenetics as we described it is largely an epistemologicalenterprise, biological systematics as a whole is filled with importantand controversial metaphysical issues which might seem to be largelyindependent of problems of inference. Whether you treat taxa astraditional natural kinds (Devitt 2008), homeostatic property clusters(Boyd 1999; Wilson, Barker, & Brigandt 2007), individuals(Ghiselin 1966, 1974; Hull 1976, 1978; Sober 1980; Ereshefsky 1991;Haber 2016a) or something else (Slater 2013, 2014) will typicallycarry little to no impact on debates about phylogeneticinference—especially as those played out in the philosophicalliterature.
Yet, early phylogeneticists certainly saw metaphysical implications oftheir views, and this impacted how they approach the problem ofreconstructing phylogeny. Simply put, if you think that taxa arehistorical entities, and that the aim of phylogeny is thereconstruction of their histories, the task is very different than ifyou take the aim to be to find ahistorical similarities or sharedessential characters (Hull 1970; Griffiths 1974). Though given theubiquitous adoption of the phylogenetic project (in a broad sense),debates over how metaphysics ought to impact inferential methodscontinues in some areas of phylogenetics.
Do competing metaphysical views in systematics impact phylogeneticinference? Yes, though only obliquely and indirectly, and it’svery important to take care not to conflate issues of classificationwith the problem of reconstructing phylogenies—even when theyoverlap. Let’s take a very brief look at the former, followed bysome commentary on the oblique and indirect way it may overlap withphylogenetic inference.
Ereshefsky (2001) argues that a commitment to phylogenetic principlesentails a reformation of the practice and norms around biologicalclassification. Specifically, he recommends leaving behind thetraditional Linnaean classification naming practices in favor of aphylogenetically informed practice. This follows a tradition stemmingback to the very earliest phylogenetic literature that sought toreform biological classification around the principle of monophyly(e.g., Hennig 1966; Griffiths 1976; Wiley 1981). For manyphylogeneticists this goal continues to this day—most recentlyaround the development of the PhyloCode (seeOther Internet Resources; de Queiroz & Gauthier 1990, 1992, 1994; Cantino & de Queiroz2020). Though structural reforms to wholly replace Linnaean taxonomyhave struggled to gain wide-scale traction and adoption, taxonomistshave largely adopted monophyly as a guiding principle inclassification and systematics—even in the context of so-calledtraditional taxonomy (though microbial systematics has proven moreresistant; see§3.1).
Of course,how biologists reconstruct a phylogeny andidentify monophyletic groups is a matter of taxonomic freedom on thePhyloCode and other phylogenetic classifications. That’s just tosay that phylogenetic inference will operate largely independently ofthese metaphysical and classificatory issues—except insofar asthese classifications inform biologists aboutwhat it is thatthey are aiming to reconstruct a history of in the first place. Haber(2019) provides a framing for this challenge, arguing that the unitsof phylogeny are the units of divergence and diversification. WhereEreshefsky (2001) and other advocates of phylogenetic classificationprovide a specific way of cashing out those units, Haber describesthis as a rich, general research project, similar to other“unit” problems in philosophy of biology. In much the sameway that how we cash out, say, the units of selection may impact howwe study or draw inferences about evolutionary processes, how weidentify the units of divergence and diversification will impact howwe draw inferences in phylogenetics. (See de Queiroz 2007 for relatedfoundational issues.)
Padian & Horner (2002) offer a coarse-grained example of how ametaphysical commitment overlaps with issues of phylogenetic inferencein their defense of treating Aves as the sole living branch ofdinosaurs. In this case, they identify “transformationist”thinking against typological thinking. In the former, the identity oftaxa is associated with historical continuity, and untethered from theexpression of any intrinsic characters. Thus, the transformation ofancient theropod dinosaurs into the living clade of birds meansthat birdsare dinosaurs, and that this ought to be reflectedin our phylogenetic reconstructions. In contrast, they argue, atypological account will treat birds as distinct from dinosaurs, andrisks a phylogenetic reconstruction that misrepresents the historicalrelationship of these taxa. Moreover, they argue thattransformationist and typological approaches will differentiallycategorize features of avian and ancient theropods as evidence anddata; what even counts as inferentially relevant is itself at stake.Padian and Horner, here, are identifying metaphysical commitments fromphylogenetics to argue how paleontologists ought to reconstructevolutionary history, what counts as evidence, and what kinds ofevidence are important for drawing those inferences, i.e., howcharacters are coded is treated as a methodological dispute entailedby theoretical commitments. (See Haber 2016b inOther Internet Resources for further discussion.)
3. Looking Ahead: New Challenges and Opportunities
We began by noting that a phylogeny reconstructs evolutionary historyand that different types of entities have an evolutionaryhistory—for example, genes, organisms, populations, and speciesall have genealogical histories. Assume that we are trying toreconstruct the history of species or higher taxa (indeed,“phylogeny” itself is sometimes defined this way, e.g.,Wiley 1981; S. Edwards 2009). Standard methods such as parsimony,likelihood, and bayesian methods all directly infer the history ofcharacters. So what is the connection between the history of thesecharacters and the phylogeny of the species? The unstated assumptionis that the histories of these characters are the same and line upwith the species history. We conclude this entry by acknowledging thatthis unstated assumption is often violated and we consider what effectthis has on the theory and practice of phylogenetic inference. As wewill see, examining this unstated assumption forces us to directlyface the question of what a phylogeny actually represents and thuswhat “phylogenetic inference” even means.
In order to facilitate an entry point into this topic we treatedphylogenetic trees and the histories they represent as simplebifurcating entities, i.e., as straight-forward branch-and-nodestructures. Yet, this is an idealization, and actual phylogenetichistories are typically more complex than this. Branches can fusetogether, and branching events turn out to be more complex than theunidimensional instantaneous point a node might suggest.
As philosophers of science are well aware, idealization in science isboth common and serves important purposes (e.g., Cartwright 1983;McMullin 1985; Wimsatt 2007; Weisberg 2007). Yet, it can be fruitfulto pause and consider if some of these idealizations and simplifyingassumptions need to be re-examined, are hindering studies, orgenerating other road blocks. Indeed, that’s precisely what hashappened recently in phylogenetics. Re-examining the utility ofsimplifying assumptions in phylogenetics has come both from outsidethe field (particularly from microbiology) and internally (asphylogeneticists began to appreciate what might be gained fromincorporating more complexity). This is (or, at least, can be) a goodand healthy thing for a science. It puts pressure on scientists andphilosophers to further articulate the central commitments of ascience, and to consider how to account for new challenges. Challengescan also push a science to adapt and revise those central commitmentsas new evidence, theories, and tools are discovered or developed, orto revisit the utility of underlying simplifying assumptions andidealizations. This, in turn, helps philosophers of science understandhow science can advance. (Haber 2019 describes this in the context ofphylogenetics asproductive disruptions.)
Revisiting the utility of idealizing away the complexity ofphylogenetic histories has generated both hard challenges and newopportunities for phylogeneticists moving forward (Haber 2012, 2019;Velasco 2012). Here we provide a brief overview of what relaxing theseassumptions amounts to, and what kinds of complexity we areacknowledging when these assumptions no longer hold. Following this,we briefly describe how this impacts the central commitments andproject of phylogenetics.
3.1 Reticulation & Discordance
In the previous sections we treated phylogenetic trees asstraight-forwardly bifurcating models, regularly referring back toFigure 1. Branches represent ancestral lineages, and nodes represent lineagesplits. Yet, this view of phylogenetics is also highly idealized,reflecting at least three simplifying assumptions: (1) that lineagessplit but do not fuse; (2) that changes on one lineage do not affectother lineages, and (3) that splits are singular, simple events. Eachof these assumptions turn out to be false.
In order to think about how lineages split or fuse, we have to firstanswer the question of what a lineage represents. In the case of aphylogeny where the tips are species, the natural answer is that thelineages are tracking species through time and the nodes representspeciation events. But consider the assumption that lineages split butdo not fuse. For many readers, hybridization may be the most obviouscounter-example. After all, hybridization is relatively familiar,especially for cases involving sexually reproducing charismaticmega-fauna and, even more frequently, plants.
In the case of speciation by hybridization, it is clear that entirelineages can fuse. Exactly how often this occurs is disputed and ofcourse will depend on how you define species in the first place. Butit is a widespread occurrence in plants and is known to happen inanimals as well (Mallet 2007; Schumer, Rosenthal, & Andolfatto2014).
While most occurrences of hybridization do not lead to new species,many often lead to introgression–that is, the introduction ofgenetic information from one lineage to another. If we model this asone lineage connecting to another, we violate the assumption that aphylogeny is a strictly branching tree diagram. But if we instead donot depict these lineages as directly connected, then we violateanother background assumption in phylogenetics, namely, that changesalong one branch do not affect any other branches. In either case, wenow have a situation where the branching history of some characters isnot the same as the history of others or of the history of the speciesas a whole. In the case of genetic history, this is known asgenealogical discordance.
While hybridization among sexually reproducing species alreadyprovides good counter-examples to our assumptions about theindependence of lineages, the bigger challenges have come frommicrobiology . Morespecifically, what we find in microbial genomics are high levels ofhorizontal (or lateral) gene transfer (HGT or LGT). For readersunfamiliar with microbial genomics, this can be thought of as themicrobial analogue to hybridization, though there are importantqualitative and quantitative differences.
Qualitatively, HGT is not a mode of reproduction, though it doesinvolve the transfer of genomic parts between microbes. Unlike inreproduction, the genomic transfer is not (typically) between parentand offspring microbes, but adjacent ones. Roughly, when microbes makecontact, they can exchange parts of their genome, which means that thehistory of these genomic parts may be distinct from the history of themicrobial organisms in which we find those genetic units. HGT can behighly advantageous for microbes, and genomic parts can quickly spreadthrough microbial populations, especially if they carry beneficialcharacteristics. Quantitatively, the rate of HGT in microbes isenormously high and commonplace, especially relative to hybridizationin, say, mammalian species.
The upshot of widespread HGT is that microbial genes will frequentlyhave different histories, and genetic parts will sometimes flow fromone lineage to very distantly related lineages. Microbialists andphilosophers of biology have taken up the question of whetherreticulation in microbial histories is enough to entirely undercut theutility of phylogenetics in microbial genomics, and how this hasimpacted the utility of thetree of life metaphor moregenerally (e.g., Martin 1999; Doolittle 1999; Velasco 2013 see too thespecial issue ofBiology & Philosophy [O’Malley,Martin, & Dupré 2010]).
The final simplifying assumption treats lineage splits as singularevents that wholly separate a lineage into two distinct new branchesthat we can cleanly track. When we think about the process ofspeciation, it is clear that speciation and lineage separation moregenerally is often not instantaneous (de Queiroz 1998; Harrison 1998).What is less clear is how this matters to phylogenetics. Thinkingcarefully about lineage splits and the internal structure of lineagesforces us to examine the nature and relationships of the multilevelorganization of the lineages we track (Haber 2012, 2019). Moreinclusive lineages contain less-inclusive lineages as parts,reflecting the hierarchical organization found in biology–afeature of lineages well appreciated by early phylogeneticists, e.g.,Hennig 1966; Griffiths 1974). Species trees, for example, contain genetrees within their branches (Maddison 1997).
When individual gene histories are incongruent with each other or withtheir containing species trees we have genealogical discordance. Wehave already seen how hybridization and horizontal gene transfer cancause discordance, but as systematics has become increasinglyconnected with population genetics, another source of discordance hasloomed large—incomplete lineage sorting. Any two copies of agene in a population will share a common ancestor in the past calledtheir point of coalescence. If two gene lineages in the same specieslineage fail to coalesce within that species branch (that is, theirpoint of coalescence is earlier than than the most recent speciationor lineage branching) we have what is calledincomplete lineagesorting (ILS). In a case of incomplete lineage sorting it ispossible that one of the gene lineages first coalesces with a genelineage from another species and thus is a case of genealogicaldiscordance. This generates what Avise and Robinson (2008) have termedhemiplasies, i.e., homologous characters whose phylogenetichistories are topologically discordant with higher level phylogeniesdue to ILS. Maddison (1997) provides a very readable introduction tothe relationship between gene trees and species trees and the problemsof genealogical discordance while Degnan and Rosenberg (2009) is aslightly more advanced introduction to the biology of incompletelineage sorting and its affects on phylogenetic inference.
To summarize, we first discussed phylogenetic inference as it wastypically done prior to around the 1990s. First, infer the history ofa number of characters (for example, genes). Each of these historiesis a particular, branching tree. Now assume that these histories arethe same as the species history. It is this assumption that allowsmethods such as parsimony or likelihood to evaluate competinghypotheses. The species history is then also a branching tree eventhough we know that in general, this is a kind of idealizationassuming that (1) that lineages split but do not fuse, (2) thatinformation is not transferred between lineages , and (3) that splitsare singular, simple events. Each of these assumptions turn out to befalse.
3.2 Impacts on Phylogenetic Inference
The reticulation and genealogical discordance generated byhybridization, HGT, ILS, and other complexities stemming from thestructure and processes involving biological lineages have,historically, been obscured by idealizing assumptions inphylogenetics. In many cases treating phylogenetic trees as idealizedmodels of genealogical history is useful and appropriate for theresearch question at hand.Figure 1 offers a good example of this. This reflects how the level ofresolution relative to our research questions can drive the way we usephylogenetic tools (O’Hara 1993).
But what happens when these idealizing assumptions are criticallyinterrogated? First, the fact that different entities have different,often incompatible, genealogical histories brings into question thevery nature of phylogeny. One line of thought is that a phylogenyshould directly track genetic history rather than the history ofspeciation. As Maddison (1995: 285) puts it,
one possible interpretation of a species phylogeny is that it depictsthe lines by which genetic information was passed on and nothingmore.
Baum and Shaw (1995), Baum (2007), and Velasco (2010, 2019) representa series of papers devoted to developing this “concordancetree” idea. Baum (2009) incorporates this line of thought aboutphylogenetic history into the debate over the nature of species andother taxa.
Philosophers have paid particular attention to reticulation in thecontext of microbial genomics, and for good reason (O’Malley etal. 2010 is a notable special edition dedicated to this topic and is auseful resource). These are often framed as challenges to thephylogenetic approach itself, or, in other cases, how and whether thetools of phylogenetics apply to microbial genomics (Martin 1999;Doolittle 1999). The breadth of HGT in microbial systems has also ledto challenges to the more conceptual and metaphysical commitmentsabout species and other phylogenetic units (Franklin 2007).
However, the majority of work in phylogenetics today continues withthe same goals as before, but with biologists using better and morepowerful tools for incorporating these complexities into theirphylogenetic models. Haber (2019) offers a philosophical treatment ofhow reticulation and genealogical discordance introduces bothchallenges and opportunities in the context of phylogenetic inference,asking whether the branches of phylogenetic trees are “toothin” and obscure relevant details found in the internalstructure of those lineages. There are a lot of good reasonsphylogeneticists might aim to include reticulate structure in theirphylogenetic reconstructions. Among other things, it means treatinggenealogical discordance as data that can be mined, as opposed tonoise that needs to be filtered out (Haber 2012, 2019). Philosophershave begun paying more attention to ways that reticulation anddiscordance have impacted phylogenetic inference beyond microbialgenomics. Rather than coming as challenges from cognate fields, theseinclude ways that practicing phylogeneticists have sought to accountfor and accommodate what we have come to understand about reticulationand discordance (see Haber & Molter 2019 for a special issuededicated to this topic).
Population biologists were among the first to operationalize methodsfor extracting information about the history of lineages from theinternal structure of those lineages, i.e., by examining the structureof individuals in a populations, population biologists developedmethods for inferring historical coalescent points of thoseindividuals (Hudson 1983; Kingman 1982, 2000; Tajima 1983). Thesemethods exploit the fact that after a lineage splits it takes time forthat split to be wholly reflected in the kinship relations of theparts of those lineages. Or, to put it in more familiar terms, ittakes time for the parts of a lineage to all be more closely relatedto each other than any are to parts in other lineages These approachescan be viewed as the continued development of methods in the largercontext of the close ties between the application of statistical toolsand molecular biology (§1.2.3).
Maddison (1997) represents a more recent launching point for a newappreciation of the role that the internal structure of lineages canplay in phylogenetic inference. Briefly, the continued advancement ofmolecular and computational techniques provided the opportunity toproduce ever-finely resolved phylogenies of the systems of interest.What quickly became apparent was a confirmation of what is onlyimplicit in Hennig (1966): that the hierarchical structure of taxa mayresult in discordant phylogenies at different levels (or even withinlevels) of the biological hierarchy. As an example, Maddison describeshow the phylogenies of individual genes or nucleotides may displaydifferent phylogenetic topographies from the organisms or populationsin which we sampled those genes. This is not a result of error orhomoplasy, but of genuine phylogenetic discordance between gene levellineages and the higher-level lineages in which they reside.
Maddison (1997) is also directly about how we ought to incorporatewhat we learn about the internal structure of lineages into our modelsof phylogenetic inference, and how we might build on the earlier workon coalescent models. Maddison is among the first to deeply appreciatehow impactful the then-new work on genomics might be forphylogenetics, both in the ways that discordance will present seriouschallenges and opportunities for phylogenetic inference. This hashelped spur new developments in this field of phylogenetics, (e.g.,Maddison & Knowles 2006; Degnan & Rosenberg 2006, 2009; Baum2007; Nakhleh 2013; Hahn & Nakhleh 2016; Mallet, Besansky, &Hahn 2016).
Haber (2019) offers two reasons why these new developments inphylogenetic inference should be of interest to philosophers. First,he offers a real-time example of scientists shifting from treating afeature of a system as noise to recognizing it as powerful evidence.Pease et al. (2016) is a prime example of this. They use some of thenew tools developed to draw phylogenetic inference in highlydiscordant systems to offer a reconstruction of the tomato clade.Where earlier analyses regarded the system as too noisy to pull out astrong phylogenetic signal, these new models permitted areconstruction that recognized specific patterns of discordance. Thisrepresents a substantial advancement in the level of resolutionphylogeneticists are able to reconstruct and identify.
Second, the new attitudes toward discordance and reticulation inphylogenetics provide an example of a community of scientists shiftingfrom one set of commitments to another. Haber (2019) offers a specificexample. He describes one of the entrenched core commitments inphylogenetic methodology as being the goal of “resolving”competing phylogenetic trees (Felsenstein 1988). That is, in caseswhere different analyses or data are generating conflictingphylogenetic trees, the goal should be to resolve this conflict byidentifying which of those trees is best supported. On a model thatrecognizes genealogical discordance, though, conflicting trees mayaccurately reflect a reticulate or discordant phylogenetic history,e.g., if different data strongly support different phylogenetic trees,one possible explanation is that we have discovered an instance ofdiscordance. Nakhleh (2013) recommends that this entails thatphylogeneticists must now also consider whether conflicting trees maybe “reconciled” rather than resolved. Haber (2019) arguesthat this is an example of what he calls a “productivedisruption”, which is one way that science might advance.
The conceptual and epistemological impacts of discordance onphylogenetic inference come together in recent debates over Sukumaranand Knowles’ (2017) work and commentary on the multispeciescoalescent (Rannala & Yang 2003; Yang & Rannala 2010). Veryroughly, the multispecies coalescent extends the principle of thecoalescent from population genetics to phylogenetics. In populationgenetics, a gene can be sampled from multiple individuals in a singlepopulation; coalescent theory provides a way to model the ancestralhistory of that gene to a common ancestor, i.e., where that historycoalesces. The multispecies coalescent does something similar, butwith samples from multiple populations (or even species). These andother techniques were developed in response to the challenge posed byMaddison (1997) to develop methods that recognize speciation as anextended process. Roughly, the lineages of genes, individualorganisms, populations, and species within a taxa can all “comeapart”, or have different phylogenetic histories, and display“a fractal hierarchy of divergences” (Sukumaran &Knowles 2017: 1607). Multispecies coalescent techniques help revealthe fine-grained structure of complex multi-level lineages (Haber2012), which has generated discussion on how these internal lineagestructures relate to species delimitation and other phylogeneticinferences (e.g., de Queiroz 2007; Knowles & Carstens 2007; Haber2012; Carstens, Pelletier, Reid, & Satler 2013; Hahn & Nakhleh2016; Mallet et al. 2016; Velasco 2019).
Sukumaran and Knowles (2017) provide a good example of the discussionsover what kinds of inferences are licensed by the multispeciescoalescent, and the way that metaphysical concerns overlap withepistemological ones. They focus on what kinds of inferences may bedrawn from multispecies coalescent methods, arguing that it is amistake to simply treat the units identified by these methods asspecies, when they may turn out to be population-level structuresinstead (i.e., populations of a species, rather than distinctspecies). They seek to highlight that mistaking population structuresfor species (or too quickly drawing the inference of the latter fromthe former, without further evidence), can undermine precisely thesorts of inferences that phylogenetics seeks to license byartificially and unjustifiably inflating species counts:
Specifically, all fields that rely on species as units of analysis,from conservation biology to studies of macroevolutionary dynamics,will be impacted by inflated estimates of the number of species,especially as genomic resources provide unprecedented power fordetecting increasingly finer-scaled genetic structure under themultispecies coalescent. (Sukumaran & Knowles 2017: 1607)
This may be framed as a claim about the units of divergence anddiversification being identified by these multispecies coalescentmethods (a metaphysical claim), and the sorts of inferences that maybe drawn about those units from these methods (an epistemologicalclaim) (Haber 2019).
To be clear, the relation between the metaphysics and epistemology isnot direct, but oblique, yet important in regard to the inferenceslicensed by phylogenetic methods. Sukumaran and Knowles (2017), forexample, conclude by issuing a plea for phylogeneticists to exercise amore careful stance towards relying on genomic data alone, and inusing multispecies coalescent as a straight-forward way to identifyspecies. They instead propose a more cautious inferential stance,identifying the units identified by the multispecies coalescent aspopulation-level structures (which may or may not correspond tospecies delimitation). This is merely an example of the rich sorts ofdiscussions around good scientific inference that is available tophilosophers willing to dive into this literature.
Bibliography
- Albert, Victor A., 2006,Parsimony, Phylogeny, andGenomics, Oxford: Oxford University Press.doi:10.1093/acprof:oso/9780199297306.001.0001
- Alfaro, Michael E. and Mark T. Holder, 2006, “The Posteriorand the Prior in Bayesian Phylogenetics”,Annual Review ofEcology, Evolution, and Systematics, 37: 19–42.doi:10.1146/annurev.ecolsys.37.091305.110021
- Amemiya, Chris T., Jessica Alföldi, Alison P. Lee, ShaohuaFan, Hervé Philippe, Iain MacCallum, Ingo Braasch, TerezaManousaki, Igor Schneider, Nicolas Rohner, Chris Organ, DomitilleChalopin, Jeramiah J. Smith, Mark Robinson, Rosemary A. Dorrington,Marco Gerdol, Bronwen Aken, Maria Assunta Biscotti, Marco Barucca,Denis Baurain, et al., 2013, “The African Coelacanth GenomeProvides Insights into Tetrapod Evolution”,Nature,496(7445): 311–316. doi:10.1038/nature12027
- Andreasen, Robin O., 1998, “A New Perspective on the RaceDebate”,The British Journal for the Philosophy ofScience, 49(2): 199–225. doi:10.1093/bjps/49.2.199
- –––, 2000, “Race: Biological Reality orSocial Construct?”,Philosophy of Science, 67:S653–S666. doi:10.1086/392853
- Ané, Cécile, Bret Larget, David A. Baum, Stacey D.Smith, and Antonis Rokas, 2007, “Bayesian Estimation ofConcordance among Gene Trees”,Molecular Biology andEvolution, 24(2): 412–426. doi:10.1093/molbev/msl170
- Ankeny, Rachel, Hasok Chang, Marcel Boumans, and Mieke Boon, 2011,“Introduction: Philosophy of Science in Practice”,European Journal for Philosophy of Science, 1(3):303–307. doi:10.1007/s13194-011-0036-4
- Archibald, Jenny K., Mark E. Mort, and Daniel J. Crawford, 2003,“Bayesian Inference of Phylogeny: A Non-Technical Primer”,TAXON, 52(2): 187–191. doi:10.2307/3647388
- Avise, John C. and Terence J. Robinson, 2008, “Hemiplasy: ANew Term in the Lexicon of Phylogenetics”,SystematicBiology, 57(3): 503–507. doi:10.1080/10635150802164587
- Baldauf, S. L., 2003, “The Deep Roots of Eukaryotes”,Science, 300(5626): 1703–1706.doi:10.1126/science.1085544
- Baum, David A., 2007, “Concordance Trees, ConcordanceFactors, and the Exploration of Reticulate Genealogy”,TAXON, 56(2): 417–426. doi:10.1002/tax.562013
- –––, 2008, “Reading a Phylogenetic Tree:The Meaning of Monophyletic Groups”,Nature Education,1(1): 190.
- –––, 2009, “Species as Ranked Taxa”,Systematic Biology, 58(1): 74–86.doi:10.1093/sysbio/syp011
- Baum, David A. and Kerry L. Shaw, 1995, “GenealogicalPerspectives of the Species Problem”, in Hoch and Stephenson1995: 289–303,
- Baum, David A. and Stacey D. Smith, 2013,Tree Thinking: AnIntroduction to Phylogenetic Biology, Greenwood Village, CO:Roberts and co.
- Baum, David A., Stacey DeWitt Smith, and Samuel S. S. Donovan,2005, “The Tree-Thinking Challenge”,Science,310(5750): 979–980. doi:10.1126/science.1117727
- Beatty, John, 1982, “Classes and Cladists”,Systematic Biology, 31(1): 25–34.doi:10.1093/sysbio/31.1.25
- Bockmann, Flávio A., Marcelo R. De Carvalho, and Murilo DeCarvalho, 2013, “The Salmon, the Lungfish (or the Coelacanth)and the Cow: A Revival?”,Zootaxa, 3750(3):265–276. doi:10.11646/zootaxa.3750.3.6
- Bowler, Peter J., 1996,Life’s Splendid Drama:Evolutionary Biology and the Reconstruction of Life’s Ancestry,1860–1940, Chicago, IL: University of Chicago Press.
- Boyd, Richard, 1999, “Homeostasis, Species and HigherTaxa”, inSpecies: New Interdisciplinary Essays, RobertA. Wilson (ed.), Cambridge, MA: MIT Press, pp. 141–185.
- Brady, Ronald H., 1982, “Theoretical Issues and‘Pattern Cladistics’”,Systematic Biology,31(3): 286–291. doi:10.1093/sysbio/31.3.286
- Brower, Andrew V.Z., 2000, “Evolution is Not a NecessaryAssumption of Cladistics”,Cladistics,16(1):143–154. doi:10.1111/j.1096-0031.2000.tb00351.x
- –––, 2002, “Cladistics, Phylogeny,Evidence and Explanation—a Reply to Lee”,ZoologicaScripta, 31(2): 221–223.doi:10.1046/j.1463-6409.2002.00102.x
- –––, 2016, “Are We All Cladists?”,inThe Future of Phylogenetic Systematics: The Legacy of WilliHennig, David Williams, Michael Schmitt, and Quentin Wheeler(eds.), (Systematics Association Special Volume Series 86), Cambridge:Cambridge University Press, 88–114.doi:10.1017/CBO9781316338797.007
- –––, 2018a, “Fifty Shades ofCladism”,Biology & Philosophy, 33(1): 8.doi:10.1007/s10539-018-9622-6
- –––, 2018b, “Statistical Consistency andPhylogenetic Inference: A Brief Review”,Cladistics,34(5): 562–567. doi:10.1111/cla.12216
- –––, 2019, “Background Knowledge: TheAssumptions of Pattern Cladistics”,Cladistics, 35(6):717-731. doi:10.1111/cla.12379
- Brower, Andrew V.Z. and Randall T. Schuh, 2021,BiologicalSystematics: Principles and Applications, Ithaca, NY: CornellUniversity Press.
- Camin, Joseph H. and Robert R. Sokal, 1965, “A Method forDeducing Branching Sequences in Phylogeny”,Evolution,19(3): 311–326. doi:10.1111/j.1558-5646.1965.tb01722.x
- Cantino, Philip D. and Kevin de Queiroz, 2020,“International Code of Phylogenetic Nomenclature,Version 6”, [Cantion & de Queiroz 2020 available online].
- Carstens, Bryan C., Tara A. Pelletier, Noah M. Reid, and Jordan D.Satler, 2013, “How to Fail at Species Delimitation”,Molecular Ecology, 22(17): 4369–4383.doi:10.1111/mec.12413
- Carpenter, James M., 1987, “Cladistics of Cladists”,Cladistics, 3(4): 363–375.doi:10.1111/j.1096-0031.1987.tb00899.x
- Cartwright, Nancy, 1983,How the Laws of Physics Lie,Oxford: Oxford University Press. doi:10.1093/0198247044.001.0001
- Cavalli-Sforza, L. L. and A. W. Edwards, 1967, “PhylogeneticAnalysis. Models and Estimation Procedures”,AmericanJournal of Human Genetics, 19(3 Pt 1): 233–257. [Cavalli-Sforza and Edwards 1967 available online]
- Chang, Joseph T., 1996, “Full Reconstruction of MarkovModels on Evolutionary Trees: Identifiability and Consistency”,Mathematical Biosciences, 137(1): 51–73.doi:10.1016/S0025-5564(96)00075-2
- Chor, Benny and Tamir Tuller, 2005, “Maximum Likelihood ofEvolutionary Trees: Hardness and Approximation”,Bioinformatics, 21(suppl 1), Oxford University Press:i97–i106. doi:10.1093/bioinformatics/bti1027
- Ciccarelli, Francesca D., Tobias Doerks, Christian von Mering,Christopher J. Creevey, Berend Snel, and Peer Bork, 2006,“Toward Automatic Reconstruction of a Highly Resolved Tree ofLife”,Science, 311(5765): 1283–1287.doi:10.1126/science.1123061
- Cleland, Carol E., 2011, “Prediction and Explanation inHistorical Natural Science”,The British Journal for thePhilosophy of Science, 62(3): 551–582.doi:10.1093/bjps/axq024
- Crawford, Nicholas G., James F. Parham, Anna B. Sellas, Brant C.Faircloth, Travis C. Glenn, Theodore J. Papenfuss, James B. Henderson,Madison H. Hansen, and W. Brian Simison, 2015, “A PhylogenomicAnalysis of Turtles”,Molecular Phylogenetics andEvolution, 83: 250–257.doi:10.1016/j.ympev.2014.10.021
- Currie, Adrian, 2018,Rock, Bone, and Ruin: AnOptimist’s Guide to the Historical Sciences, Boston, MA:MIT Press.
- –––, 2019,Scientific Knowledge and the DeepPast: History Matters, Cambridge: Cambridge University Press.doi:10.1017/9781108582490
- Darwin, Charles, 1859,On the Origin of Species, London:John Murray.
- Degnan, James H. and Noah A. Rosenberg, 2006, “Discordanceof Species Trees with Their Most Likely Gene Trees”,PLoSGenetics, 2(5): e68. doi:10.1371/journal.pgen.0020068
- –––, 2009, “Gene Tree Discordance,Phylogenetic Inference and the Multispecies Coalescent”,Trends in Ecology & Evolution, 24(6): 332–340.doi:10.1016/j.tree.2009.01.009
- de Queiroz, Kevin, 1998, “The General Lineage Concept ofSpecies, Species Criteria, and the Process of Speciation: A ConceptualUnification and Terminological Recommendations”, in Howard andBerlocher 1998: 57–75.
- –––, 2007, “Species Concepts and SpeciesDelimitation”,Systematic Biology, 56(6):879–886. doi:10.1080/10635150701701083
- de Queiroz, Kevin and Jacques Gauthier, 1990, “Phylogeny asa Central Principle in Taxonomy: Phylogenetic Definitions of TaxonNames”,Systematic Zoology, 39(4): 307–322.doi:10.2307/2992353
- –––, 1992, “Phylogenetic Taxonomy”,Annual Review of Ecology and Systematics, 23(1):449–480. doi:10.1146/annurev.es.23.110192.002313
- –––, 1994, “Toward a Phylogenetic Systemof Biological Nomenclature”,Trends in Ecology &Evolution, 9(1): 27–31.doi:10.1016/0169-5347(94)90231-3
- de Queiroz, Kevin and Steven Poe, 2001, “Philosophy andPhylogenetic Inference: A Comparison of Likelihood and ParsimonyMethods in the Context of Karl Popper’s Writings onCorroboration”,Systematic Biology, 50(3):305–321. doi:10.1080/106351501300317941
- –––, 2003, “Failed Refutations: FurtherComments on Parsimony and Likelihood Methods and Their Relationship toPopper’s Degree of Corroboration”,SystematicBiology, 52(3): 352–367. doi:10.1080/10635150309324
- Devitt, Michael, 2008, “Resurrecting BiologicalEssentialism”,Philosophy of Science, 75(3):344–382. doi:10.1086/593566
- Dietrich, Michael R., 1994, “The Origins of the NeutralTheory of Molecular Evolution”,Journal of the History ofBiology, 27(1): 21–59. doi:10.1007/BF01058626
- –––, 1998, “Paradox and Persuasion:Negotiating the Place of Molecular Evolution within EvolutionaryBiology”,Journal of the History of Biology, 31(1):85–111. doi:10.1023/A:1004257523100
- Donoghue, Michael J., 1992, “Homology”, inKeywords in Evolutionary Biology, Evelyn Fox Keller andElisabeth A. Lloyd (eds.), Cambridge, MA: Harvard University Press,170–179.
- Doolittle, W. Ford, 1999, “Phylogenetic Classification andthe Universal Tree”,Science, 284(5423):2124–2128. doi:10.1126/science.284.5423.2124
- Dunn, Casey W., Gonzalo Giribet, Gregory D. Edgecombe, and AndreasHejnol, 2014, “Animal Phylogeny and Its EvolutionaryImplications”,Annual Review of Ecology, Evolution, andSystematics, 45: 371–395.doi:10.1146/annurev-ecolsys-120213-091627
- Duret, Laurent, 2008, “Neutral Theory: The Null Hypothesisof Molecular Evolution”,Nature Education, 1(1):218.
- Editors, 2016, “Editorial”,Cladistics,32(1): 1. doi:10.1111/cla.12148
- Edwards, Anthony W. F., 1970, “Estimation of the BranchPoints of a Branching Diffusion Process”,Journal of theRoyal Statistical Society: Series B (Methodological), 32(2):155–164. doi:10.1111/j.2517-6161.1970.tb00828.x
- –––, 1972,Likelihood: An Account of theStatistical Concept of Likelihood and Its Application to ScientificInference, Cambridge: Cambridge University Press.
- –––, 1996, “The Origin and EarlyDevelopment of the Method of Minimum Evolution for the Reconstructionof Phylogenetic Trees”,Systematic Biology, 45(1):79–91. doi:10.1093/sysbio/45.1.79
- Edwards, Scott V., 2009, “Is a New and General Theory ofMolecular Systematics Emerging?”,Evolution, 63(1):1–19. doi:10.1111/j.1558-5646.2008.00549.x
- Eldredge, Niles and Joel Cracraft, 1980,Phylogenetic Patternsand the Evolutionary Process, New York: Columbia UniversityPress.
- Ereshefsky, Marc, 1991, “Species, Higher Taxa, and the Unitsof Evolution”,Philosophy of Science, 58(1):84–101. doi:10.1086/289600
- –––, 2001,The Poverty of the LinnaeanHierarchy, Cambridge: Cambridge University Press.doi:10.1017/CBO9780511498459
- –––, 2012, “Homology Thinking”,Biology and Philosophy, 27(3): 381–400.doi:10.1007/s10539-012-9313-7
- Fagan, Melinda Bonnie, 2013,Philosophy of Stem Cell Biology:Knowledge in Flesh and Blood, New York: Palgrave Macmillan.doi:10.1057/9781137296023
- Farris, James S., 1983, “The Logical Basis of PhylogeneticAnalysis”, inAdvances in Cladistics: Proceedings of theThird Meeting of the Willi Hennig Society, Vol. 2, N. I. Platnick& V. A. Funk (eds.), New York: Columbia University Press, pp.7–36.
- –––, 1999, “Likelihood andInconsistency”,Cladistics, 15(2): 199–204.doi:10.1111/j.1096-0031.1999.tb00262.x
- Felsenstein, Joseph, 1973, “Maximum Likelihood andMinimum-Steps Methods for Estimating Evolutionary Trees from Data onDiscrete Characters”,Systematic Zoology, 22(3):240–249. doi:10.1093/sysbio/22.3.240
- –––, 1978, “Cases in Which Parsimony orCompatibility Methods Will Be Positively Misleading”,Systematic Zoology, 27(4): 401–410.doi:10.2307/2412923
- –––, 1981, “Evolutionary Trees from DNASequences: A Maximum Likelihood Approach”,Journal ofMolecular Evolution, 17(6): 368–376.doi:10.1007/BF01734359
- –––, 1983, “Parsimony in Systematics:Biological and Statistical Issues”,Annual Review of Ecologyand Systematics, 14(1): 313–333.doi:10.1146/annurev.es.14.110183.001525
- –––, 1988, “Phylogenies from MolecularSequences: Inference and Reliability”,Annual Review ofGenetics, 22(1): 521–565.doi:10.1146/annurev.ge.22.120188.002513
- –––, 2001, “The Troubled Growth ofStatistical Phylogenetics”,Systematic Biology, 50(4):465–467. doi:10.1080/10635150119297
- –––, 2004,Inferring Phylogenies,volume 2, Sunderland, MA: Sinauer Associates.
- Felsenstein, Joseph and Elliott Sober, 1986, “Parsimony andLikelihood: An Exchange”,Systematic Zoology, 35(4):617–626. doi:10.2307/2413121
- Fisher, Ronald A., 1935 [1950],Statistical Methods forResearch Workers, Edinburgh: Oliver and Boyd. Eleventh edition1950.
- –––, 1956,Statistical Methods andScientific Inference, first edition, Edinburgh: Oliver andBoyd.
- Fitch, Walter M. and Emanuel Margoliash, 1967, “Constructionof Phylogenetic Trees: A Method Based on Mutation Distances asEstimated from Cytochromec Sequences Is of GeneralApplicability”,Science, 155(3760): 279–284.doi:10.1126/science.155.3760.279
- Fitzhugh, Kirk, 2008, “Abductive Inference: Implications for‘Linnean’ and ‘Phylogenetic’ Approaches forRepresenting Biological Systematization”,EvolutionaryBiology, 35(1): 52–82. doi:10.1007/s11692-008-9015-x
- Franklin, L. R., 2007, “Bacteria, Sex, andSystematics”,Philosophy of Science, 74(1):69–95. doi:10.1086/519476
- Freudenstein, John V., 2005, “Characters, States andHomology”,Systematic Biology, 54(6): 965–973.doi:10.1080/10635150500354654
- Gaffney, Eugene S., 1979, “An Introduction to the Logic ofPhylogeny Reconstruction”, inPhylogenetic Analysis andPaleontology, Joel Cracraft and Niles Eldredge (eds), New York:Columbia University Press, pp. 79–111.
- Gardiner, B. G., P. Janvier, C. Patterson, P. L. Forey, P. H.Greenwood, R. S. Miles, and R. P. S. Jefferies, 1979, “TheSalmon, the Lungfish and the Cow: A Reply”,Nature,277(5693): 175–176. doi:10.1038/277175b0
- Ghiselin, Michael T., 1966, “On Psychologism in the Logic ofTaxonomic Controversies”,Systematic Zoology, 15(3):207–215. doi:10.2307/2411392
- –––, 1974, “A Radical Solution to theSpecies Problem”,Systematic Zoology, 23(4):536–544. doi:10.2307/2412471
- Giere, Ronald N., 1979 [1997],Understanding ScientificReasoning, New York: Holt, Rinehart, and Winston. Fourth edition,Fort Worth, TX: Harcourt Brace College Publishers, 1997.
- Goodman, Nelson, 1972, “Seven Strictures onSimilarity”, in hisProblems and Projects,Indianapolis, IN: The Bobbs-Merrill Company, pp. 437–446.
- Graham, R.L. and L.R. Foulds, 1982, “Unlikelihood ThatMinimal Phylogenies for a Realistic Biological Study Can BeConstructed in Reasonable Computational Time”,MathematicalBiosciences, 60(2): 133–142.doi:10.1016/0025-5564(82)90125-0
- Grantham, Todd, 2004, “The Role of Fossils in PhylogenyReconstruction: Why Is It so Difficult to Integrate Paleobiologicaland Neontological Evolutionary Biology?”,Biology andPhilosophy, 19(5): 687–720.doi:10.1007/s10539-005-0370-z
- Gray, Russell D. and Quentin D. Atkinson, 2003,“Language-Tree Divergence Times Support the Anatolian Theory ofIndo-European Origin”,Nature, 426(6965):435–439. doi:10.1038/nature02029
- Gray, Russell D., A. J. Drummond, and S. J. Greenhill, 2009,“Language Phylogenies Reveal Expansion Pulses and Pauses inPacific Settlement”,Science, 323(5913): 479–483.doi:10.1126/science.1166858
- Gray, Russell D. and Fiona M. Jordan, 2000, “Language TreesSupport the Express-Train Sequence of Austronesian Expansion”,Nature, 405(6790): 1052–1055. doi:10.1038/35016575
- Griesemer, James, 2000, “Development, Culture, and the Unitsof Inheritance”,Philosophy of Science, 67(S1):S348–S368. doi:10.1086/392831
- Griffiths, Graham C. D., 1974, “On the Foundations ofBiological Systematics”,Acta Biotheoretica,23(3–4): 85–131. doi:10.1007/BF01556343
- –––, 1976, “The Future of LinnaeanNomenclature”,Systematic Zoology, 25(2):168–173. doi:10.2307/2412743
- Haber, Matthew H., 2005, “On Probability and Systematics:Possibility, Probability, and Phylogenetic Inference”,Systematic Biology, 54(5): 831–841.doi:10.1080/106351591007444
- –––, 2009, “Phylogenetic Inference”,inA Companion to the Philosophy of History andHistoriography, Aviezer Tucker (ed.), (Blackwell Companions toPhilosophy 41), Oxford, UK: Wiley-Blackwell, 231–242.doi:10.1002/9781444304916.ch20
- –––, 2012, “Multilevel Lineages andMultidimensional Trees: The Levels of Lineage and PhylogenyReconstruction”,Philosophy of Science, 79(5):609–623. doi:10.1086/667849
- –––, 2016a, “The Individuality Thesis (3Ways)”,Biology & Philosophy, 31(6): 913–930.doi:10.1007/s10539-016-9548-9
- –––, 2019, “Species in the Age ofDiscordance”,Philosophy, Theory, and Practice inBiology, 11: art. 21. doi:10.3998/ptpbio.16039257.0011.021
- Haber, Matthew H. and Daniel J. Molter, 2019, “Species inthe Age of Discordance: Meeting Report and Introduction”,Philosophy, Theory, and Practice in Biology, 11: art. 12.doi:10.3998/ptpbio.16039257.0011.012
- Hagen, Joel B., 2001, “The Introduction of Computers intoSystematic Research in the United States during the 1960s”,Studies in History and Philosophy of Science Part C: Studies inHistory and Philosophy of Biological and Biomedical Sciences,32(2): 291–314. doi:10.1016/S1369-8486(01)00005-X
- Hahn, Matthew W. and Luay Nakhleh, 2016, “IrrationalExuberance for Resolved Species Trees”,Evolution,70(1): 7–17. doi:10.1111/evo.12832
- Hall, Brian K., 1994,Homology: The Hierarchical Basis ofComparative Biology, San Diego, CA: Academic Press.
- Halstead, L. B., 1978, “The Cladistic Revolution–CanIt Make the Grade?”,Nature, 276(5690): 759–760.doi:10.1038/276759a0
- Harrison, Richard G., 1998, “Linking Evolutionary Patternand Process”, in Howard and Berlocher 1998: 19–31.
- Hasegawa, Masami, Hirohisa Kishino, and Taka-aki Yano, 1985,“Dating of the Human-Ape Splitting by a Molecular Clock ofMitochondrial DNA”,Journal of Molecular Evolution,22(2): 160–174. doi:10.1007/BF02101694
- Havstad, Joyce C. and N. Adam Smith, 2019, “Fossils withFeathers and Philosophy of Science”,SystematicBiology, 68(5): 840–851. doi:10.1093/sysbio/syz010
- Heckman, Daniel S., David M. Geiser, Brooke R. Eidell, Rebecca L.Stauffer, Natalie L. Kardos, and S. Blair Hedges, 2001,“Molecular Evidence for the Early Colonization of Land by Fungiand Plants”,Science, 293(5532): 1129–1133.doi:10.1126/science.1061457
- Hennig, Willi, 1966,Phylogenetic Systematics, D. DwightDavis and Rainer Zangerl (trans.), Urbana, IL: University of IllinoisPress; this is a revised and translated version of the Germanoriginal,Grundzüge einer Theorie der phylogenetischenSystematik, Berlin: Deutscher Zentralverlag, 1950.
- –––, 1975, “‘Cladistic Analysis orCladistic Classification’: A Reply to Ernst Mayr”,Systematic Zoology, 24(2): 244–256.doi:10.1093/sysbio/24.2.244
- Hillis, David M. and James J. Bull, 1993, “An Empirical Testof Bootstrapping as a Method for Assessing Confidence in PhylogeneticAnalysis”,Systematic Biology, 42(2): 182–192.doi:10.1093/sysbio/42.2.182
- Hillis, David M., James J. Bull, Mary E. White, Marty R. Badgett,and Ian J. Molineux, 1992, “Experimental Phylogenetics:Generation of a Known Phylogeny”,Science, 255(5044):589–592. doi:10.1126/science.1736360
- Hillis, David M., John P. Huelsenbeck, and Clifford W. Cunningham,1994, “Application and Accuracy of Molecular Phylogenies”,Science, 264(5159): 671–677.doi:10.1126/science.8171318
- Hinchliff, Cody E., Stephen A. Smith, James F. Allman, J. GordonBurleigh, Ruchi Chaudhary, Lyndon M. Coghill, Keith A. Crandall,Jiabin Deng, Bryan T. Drew, Romina Gazis, Karl Gude, David S. Hibbett,Laura A. Katz, H. Dail Laughinghouse, Emily Jane McTavish, Peter E.Midford, Christopher L. Owen, Richard H. Ree, Jonathan A. Rees,Douglas E. Soltis, et al., 2015, “Synthesis of Phylogeny andTaxonomy into a Comprehensive Tree of Life”,Proceedings ofthe National Academy of Sciences, 112(41): 12764–12769.doi:10.1073/pnas.1423041112
- Hoch, Peter C. and A. G. Stephenson (eds), 1995,Experimentaland Molecular Approaches to Plant Biosystematics, St. Louis, MO:Proceedings of the Fifth International Symposium of the InternationalOrganization of Plant Biosystematists; Missouri Botanical GardenPress.
- Howard, Daniel J. and Stewart H. Berlocher (eds.), 1998,Endless Forms: Species and Speciation, New York: OxfordUniversity Press.
- Hudson, Richard R., 1983, “Properties of a Neutral AlleleModel with Intragenic Recombination”,Theoretical PopulationBiology, 23(2): 183–201.doi:10.1016/0040-5809(83)90013-8
- Huelsenbeck, John P., 1997, “Is the Felsenstein Zone a FlyTrap?”,Systematic Biology, 46(1): 69–74.doi:10.1093/sysbio/46.1.69
- Huelsenbeck, John P., Cécile Ané, Bret Larget, andFredrik Ronquist, 2008, “A Bayesian Perspective on aNon-Parsimonious Parsimony Model”,Systematic Biology,57(3): 406–419. doi:10.1080/10635150802166046
- Hug, Laura A., Brett J. Baker, Karthik Anantharaman, ChristopherT. Brown, Alexander J. Probst, Cindy J. Castelle, Cristina N.Butterfield, Alex W. Hernsdorf, Yuki Amano, Kotaro Ise, Yohey Suzuki,Natasha Dudek, David A. Relman, Kari M. Finstad, Ronald Amundson,Brian C. Thomas, and Jillian F. Banfield, 2016, “A New View ofthe Tree of Life”,Nature Microbiology, 1(5): 16048.doi:10.1038/nmicrobiol.2016.48
- Hull, David L., 1970, “Contemporary SystematicPhilosophies”,Annual Review of Ecology andSystematics, 1(1): 19–54.doi:10.1146/annurev.es.01.110170.000315
- –––, 1976, “Are Species ReallyIndividuals?”,Systematic Zoology, 25(2):174–191. doi:10.2307/2412744
- –––, 1978, “A Matter ofIndividuality”,Philosophy of Science, 45(3):335–360. doi:10.1086/288811
- –––, 1983, “Karl Popper and Plato’sMetaphor”, inAdvances in Cladistics, Vol. 2, N. I.Platnick & V. A. Funk (eds.), New York: Columbia University Press,pp. 177–189.
- –––, 1988,Science as a Process: AnEvolutionary Account of the Social and Conceptual Development ofScience, Chicago, IL: The University of Chicago Press.
- –––, 1999, “The Use and Abuse of Sir KarlPopper”,Biology and Philosophy, 14(4): 481–504.doi:10.1023/A:1006554919188
- Huxley, Julian (ed.), 1940,The New Systematics., Oxford:Clarendon Press.
- Jukes, Thomas H. and Charles R. Cantor, 1969, “Evolution ofProtein Molecules”, inMammalian Protein Metabolism,volume 3, New York: Academic Press, 21–132.doi:10.1016/B978-1-4832-3211-9.50009-7
- Kearney, Maureen, 2007, “Philosophy and Phylogenetics:Historical and Current Connections”, inThe CambridgeCompanion to the Philosophy of Biology, David L. Hull and MichaelRuse (eds.), Cambridge: Cambridge University Press, 211–232.doi:10.1017/CCOL9780521851282.011
- Kendig, Catherine, 2016, “Homologizing as Kinding”, inNatural Kinds and Classification in Scientific Practice,Catherine Kendig (ed.), London: Routledge, 106-125.
- Kenrick, Paul and Peter R. Crane, 1997, “The Origin andEarly Evolution of Plants on Land”,Nature, 389(6646):33–39. doi:10.1038/37918
- Kimura, Motoo, 1968, “Evolutionary Rate at the MolecularLevel”,Nature, 217(5129): 624–626.doi:10.1038/217624a0
- –––, 1969, “The Rate of MolecularEvolution Considered from the Standpoint of PopulationGenetics”,Proceedings of the National Academy ofSciences, 63(4): 1181–1188. doi:10.1073/pnas.63.4.1181
- –––, 1980, “A Simple Method for EstimatingEvolutionary Rates of Base Substitutions through Comparative Studiesof Nucleotide Sequences”,Journal of MolecularEvolution, 16(2): 111–120. doi:10.1007/BF01731581
- Kingman, J. F. C., 1982, “On the Genealogy of LargePopulations”,Journal of Applied Probability, 19:27–43. doi:10.2307/3213548
- –––, 2000, “Origins of the Coalescent:1974–1982”,Genetics, 156(4):1461–1463.
- Kluge, Arnold G., 2001, “Philosophical Conjectures and TheirRefutation”,Systematic Biology, 50(3): 322–330.doi:10.1080/10635150119615
- –––, 2005, “What Is the Rationale for‘Ockham’s Razor’(aka Parsimony) in PhylogeneticInference”, in Albert 2005: 15–42.
- Kluge, Arnold G. and James S. Farris, 1999, “Taxic Homology= Overall Similarity”,Cladistics, 15(2): 205–212.doi:10.1111/j.1096-0031.1999.tb00263.x
- Knowles, L. Lacey and Bryan C. Carstens, 2007, “DelimitingSpecies without Monophyletic Gene Trees”,SystematicBiology, 56(6): 887–895. doi:10.1080/10635150701701091
- Larget, Bret and Donald L. Simon, 1999, “Markov Chain MonteCarlo Algorithms for the Bayesian Analysis of PhylogeneticTrees”,Molecular Biology and Evolution, 16(6):750–759. doi:10.1093/oxfordjournals.molbev.a026160
- Lee, Michael S.Y., 2002, “Divergent Evolution, Hierarchy,and Cladistics”,Zoologica Scripta, 31(2): 217–219.doi:10.1046/j.1463-6409.2002.00101.x
- Lemey, Philippe, Marco Salemi, and Anne-Mieke Vandamme (eds.),2009,The Phylogenetic Handbook: A Practical Approach toPhylogenetic Analysis and Hypothesis Testing, second edition,Cambridge: Cambridge University Press.doi:10.1017/CBO9780511819049
- Lemieux, Jacob E., Katherine J. Siddle, Bennett M. Shaw, ChristineLoreth, Stephen F. Schaffner, Adrianne Gladden-Young, Gordon Adams,Timelia Fink, Christopher H. Tomkins-Tinch, Lydia A. Krasilnikova,Katherine C. DeRuff, Melissa Rudy, Matthew R. Bauer, Kim A. Lagerborg,Erica Normandin, Sinéad B. Chapman, Steven K. Reilly, Melis N.Anahtar, Aaron E. Lin, Amber Carter, et al., 2021, “PhylogeneticAnalysis of SARS-CoV-2 in Boston Highlights the Impact ofSuperspreading Events”,Science, 371(6529):574–575. doi:10.1126/science.abe3261
- Lewens, Tim, 2012, “Pheneticism Reconsidered”,Biology and Philosophy, 27(2), Springer Netherlands:159–177. doi:10.1007/s10539-011-9302-2
- Li, Shuying, Dennis K. Pearl, and Hani Doss, 2000,“Phylogenetic Tree Construction Using Markov Chain MonteCarlo”,Journal of the American StatisticalAssociation, 95(450): 493–508.doi:10.1080/01621459.2000.10474227
- Lipton, Peter, 2004,Inference to the Best Explanation,second edition, London: Routledge. doi:10.4324/9780203470855
- Longino, Helen E., 1990,Science as Social Knowledge: Valuesand Objectivity in Scientific Inquiry, Princeton, NJ: PrincetonUniversity Press.
- –––, 2019, “The Social Dimensions ofScientific Knowledge”, inThe Stanford Encyclopedia ofPhilosophy (Summer 2019), Edward N. Zalta (ed.). URL= <https://plato.stanford.edu/archives/sum2019/entries/scientific-knowledge-social/>
- Love, Alan C., 2014, “The Erotetic Organization ofDevelopmental Biology”, inTowards a Theory ofDevelopment, Alessandro Minelli and Thomas Pradeu (eds.), Oxford:Oxford University Press, 33–55.doi:10.1093/acprof:oso/9780199671427.003.0003
- Maddison, Wayne P., 1995, “Phylogenetic Histories Within andAmong Species”, in Hoch and Stephenson 1995: 273–287.
- –––, 1997, “Gene Trees in SpeciesTrees”,Systematic Biology, 46(3): 523–536.doi:10.1093/sysbio/46.3.523
- Maddison, Wayne P. and L. Lacey Knowles, 2006, “InferringPhylogeny Despite Incomplete Lineage Sorting”,SystematicBiology, 55(1): 21–30. doi:10.1080/10635150500354928
- Mallet, James, 2007, “Hybrid Speciation”,Nature, 446(7133): 279–283.doi:10.1038/nature05706
- Mallet, James, Nora Besansky, and Matthew W. Hahn, 2016,“How Reticulated Are Species?”,BioEssays, 38(2):140–149. doi:10.1002/bies.201500149
- Margoliash, Emanuel, 1963, “Primary Structure and Evolutionof Cytochromec”,Proceedings of the NationalAcademy of Sciences, 50(4): 672–679.doi:10.1073/pnas.50.4.672
- Martin, William, 1999, “Mosaic Bacterial Chromosomes: AChallenge En Route to a Tree of Genomes”,BioEssays,21(2): 99–104.doi:10.1002/(SICI)1521-1878(199902)21:2<99::AID-BIES3>3.0.CO;2-B
- Mau, Bob and Michael A. Newton, 1997, “PhylogeneticInference for Binary Data on Dendograms Using Markov Chain MonteCarlo”,Journal of Computational and GraphicalStatistics, 6(1): 122–131.doi:10.1080/10618600.1997.10474731
- Mayr, Ernst, 1965a, “Classification and Phylogeny”,American Zoologist, 5(1): 165–174.doi:10.1093/icb/5.1.165
- –––, 1965b, “Numerical Phenetics andTaxonomic Theory”,Systematic Zoology, 14(2):73–97. doi:10.2307/2411730
- –––, 1969,Principles of SystematicZoology, New York: MacGraw-Hill.
- –––, 1974, “Cladistic Analysis orCladistic Classification?”,Journal of ZoologicalSystematics and Evolutionary Research, 12(1): 94–128.doi:10.1111/j.1439-0469.1974.tb00160.x
- McMullin, Ernan, 1985, “Galilean Idealization”,Studies in History and Philosophy of Science Part A, 16(3):247–273. doi:10.1016/0039-3681(85)90003-2
- Mishler, Brent D., 2005, “The Logic of the Data Matrix inPhylogenetic Analysis”, in Albert 2005: 57–70.
- Morgan, Gregory J., 1998, “Emile Zuckerkandl, Linus Pauling,and the Molecular Evolutionary Clock, 1959-1965”,Journal ofthe History of Biology, 31(2): 155–178.doi:10.1023/A:1004394418084
- Nakhleh, Luay, 2013, “Computational Approaches to SpeciesPhylogeny Inference and Gene Tree Reconciliation”,Trends inEcology & Evolution, 28(12): 719–728.doi:https://doi.org/10.1016/j.tree.2013.09.004
- Neff, Nancy A., 1986, “A Rational Basis for a PrioriCharacter Weighting”,Systematic Zoology, 35(1):110–123. doi:10.1093/sysbio/35.1.110
- Neyman, Jerzy, 1952,Lectures and Conferences on MathematicalStatistics and Probability, second edition, Washington, DC:Graduate School, United States Department of Agriculture.
- –––, 1971, “Molecular Studies ofEvolution: A Source of Novel Statistical Problems”, inStatistical Decision Theory and Related Topics, Shanti S.Gupta and James Yackel (eds), New York: Academic Press,1–27.
- O’Hara, Robert J., 1988, “Homage to Clio, or, Towardan Historical Philosophy for Evolutionary Biology”,Systematic Zoology, 37(2): 142–155.doi:10.2307/2992272
- –––, 1993, “Systematic Generalization,Historical Fate, and the Species Problem”,SystematicBiology, 42(3): 231–246. doi:10.1093/sysbio/42.3.231
- O’Malley, Maureen A., William Martin, and John Dupré,2010, “The Tree of Life: Introduction to an EvolutionaryDebate”,Biology and Philosophy, 25(4): 441–453.doi:10.1007/s10539-010-9208-4
- Owen, Richard, 1843,Lectures on the Comparative Anatomy andPhysiology of the Invertebrate Animals, London: Longman, Brown,Greene, and Longmans.
- Padian, Kevin and John R. Horner, 2002, “Typology versusTransformation in the Origin of Birds”,Trends in Ecology& Evolution, 17(3): 120–124.doi:10.1016/S0169-5347(01)02409-0
- Patterson, Colin, 1982, “Classes and Cladists or Individualsand Evolution”,Systematic Biology, 31(3): 284–286.doi:10.1093/sysbio/31.3.284
- Pauling, Linus and Emile Zuckerkandl, 1963, “ChemicalPaleogenetics. Molecular ‘Restoration Studies’ of ExtinctForms of Life”,Acta Chemica Scandinavica, 17 supl.:9–16. doi:10.3891/acta.chem.scand.17s-0009
- Pearson, Christopher H., 2010, “Pattern Cladism, Homology,and Theory-Neutrality”,History and Philosophy of the LifeSciences, 32(4): 475–492.
- Pease, James B., David C. Haak, Matthew W. Hahn, and Leonie C.Moyle, 2016, “Phylogenomics Reveals Three Sources of AdaptiveVariation during a Rapid Radiation”,PLOS Biology,14(2): 1–24. doi:10.1371/journal.pbio.1002379
- Pickett, K. M. and Randle, C. P., 2005, “Strange bayesindeed: uniform topological priors imply non–uniform cladepriors”,Molecular Phylogenetics and Evolution, 34(1):203–211. doing:10.1016/j.ympev.2004.09.001
- Pinna, Mário C. C. de, 1991, “Concepts and Tests ofHomology in the Cladistic Paradigm”,Cladistics, 7(4):367–394. doi:10.1111/j.1096-0031.1991.tb00045.x
- Platnick, Norman I., 1982, “Defining Characters andEvolutionary Groups”,Systematic Biology, 31(3):282–282. doi:10.1093/sysbio/31.3.282
- Posada, David, 2008, “JModelTest: Phylogenetic ModelAveraging”,Molecular Biology and Evolution, 25(7):1253–1256. doi:10.1093/molbev/msn083
- Posada, David and Keith A. Crandall, 1998, “MODELTEST:Testing the Model of DNA Substitution”,Bioinformatics,14(9): 817–818. doi:10.1093/bioinformatics/14.9.817
- Quinn, Aleta, 2016, “Phylogenetic Inference to the BestExplanation and the Bad Lot Argument”,Synthese,193(9): 3025–3039. doi:10.1007/s11229-015-0908-9
- –––, 2017, “When Is a Cladist Not aCladist?”,Biology & Philosophy, 32(4):581–598. doi:10.1007/s10539-017-9577-z
- –––, 2019, “Diagnosing Discordance: Signalin Data, Conflict in Paradigms”,Philosophy, Theory, andPractice in Biology, 11(July): art. 017.doi:10.3998/ptpbio.16039257.0011.017
- Rannala, Bruce and Ziheng Yang, 1996, “ProbabilityDistribution of Molecular Evolutionary Trees: A New Method ofPhylogenetic Inference”,Journal of MolecularEvolution, 43(3): 304–311. doi:10.1007/BF02338839
- –––, 2003, “Bayes Estimation of SpeciesDivergence Times and Ancestral Population Sizes Using DNA Sequencesfrom Multiple Loci”,Genetics, 164(4): 1645–1656.doi:10.1093/genetics/164.4.1645
- Redelings, Benjamin D. and Marc A. Suchard, 2005, “JointBayesian Estimation of Alignment and Phylogeny”,SystematicBiology, 54(3): 401–418. doi:10.1080/10635150590947041
- Richards, Richard, 2002, “Kuhnian Values and CladisticParsimony”,Perspectives on Science, 10(1): 1–27.doi:10.1162/106361402762674780
- –––, 2003, “Character Individuation inPhylogenetic Inference”,Philosophy of Science, 70(2):264–279. doi:10.1086/375467
- Rieppel, Olivier, 2008, “Re-Writing Popper’sPhilosophy of Science for Systematics”,History andPhilosophy of the Life Sciences, 30(3–4):293–316.
- Rieppel, Olivier and Maureen Kearney, 2002,“Similarity”,Biological Journal of the LinneanSociety, 75(1): 59–82.doi:10.1046/j.1095-8312.2002.00006.x
- –––, 2007, “The Poverty of TaxonomicCharacters”,Biology & Philosophy, 22(1):95–113. doi:10.1007/s10539-006-9024-z
- Roffé, Ariel Jonathan, 2020, “Dynamic Homology andCircularity in Cladistic Analysis”,Biology &Philosophy, 35(1): art. 21. doi:10.1007/s10539-020-9737-4
- Rogers, James S., 2001, “Maximum Likelihood Estimation ofPhylogenetic Trees Is Consistent When Substitution Rates VaryAccording to the Invariable Sites plus Gamma Distribution”,Systematic Biology, 50(5): 713–722.doi:10.1080/106351501753328839
- Ronquist, Fredrik, Paul van der Mark, and John P. Huelsenbeck,2009, “Bayesian Phylogenetic Analysis Using MRBAYES”, inLemey, Salemi, and Vandamme 2009: 210–266.doi:10.1017/CBO9780511819049.009
- Salipante, Stephen J. and Marshall S. Horwitz, 2006,“Phylogenetic Fate Mapping”,Proceedings of theNational Academy of Sciences, 103(14): 5448–5453.doi:10.1073/pnas.0601265103
- Santis, Marcelo Domingos de, 2020, “Popper as a process:revisiting the appropriation of the Popperian philosophy by thecladists during the ‘systematics wars’”,Arquivos De Zoologia, 51(2): 13–20.doi:10.11606/2176-7793/2020.51.02
- Schumer, Molly, Gil G. Rosenthal, and Peter Andolfatto, 2014,“How Common Is Homoploid Hybrid Speciation?”,Evolution, 68(6): 1553–1560. doi:10.1111/evo.12399
- Scotland, Robert and R. Toby Pennington (eds.), 2000,Homologyand Systematics: Coding Characters for Phylogenetic Analysis,London: CRC Press. doi:10.1201/9781482268249
- Scotland, Robert W., Richard G. Olmstead, and Jonathan R. Bennett,2003, “Phylogeny Reconstruction: The Role of Morphology”,Systematic Biology, 52(4): 539–548.doi:10.1080/10635150309309
- Sereno, Paul C., 2005, “The Logical Basis of PhylogeneticTaxonomy”,Systematic Biology, 54(4): 595–619.doi:10.1080/106351591007453
- Siddall, Mark E., 1998, “Success of Parsimony in theFour-Taxon Case: Long-Branch Repulsion by Likelihood in the FarrisZone”,Cladistics, 14(3): 209–220.doi:10.1111/j.1096-0031.1998.tb00334.x
- Siddall, Mark E. and Arnold G. Kluge, 1997, “Probabilism andPhylogenetic Inference”,Cladistics, 13(4):313–336. doi:10.1111/j.1096-0031.1997.tb00322.x
- Siddall, Mark E. and Michael F. Whiting, 1999, “Long-BranchAbstractions”,Cladistics, 15(1): 9–24.doi:10.1111/j.1096-0031.1999.tb00391.x
- Simpson, George Gaylord, 1961,Principles of AnimalTaxonomy, New York: Columbia University Press.
- Slater, Matthew H., 2013,Are Species Real? An Essay on theMetaphysics of Species, (New Directions in the Philosophy ofScience), New York: Palgrave Macmillan. doi:10.1057/9780230393233
- –––, 2015, “Natural Kindness”,The British Journal for the Philosophy of Science, 66(2):375–411. doi:10.1093/bjps/axt033
- Sneath, Peter H. A. and Robert R. Sokal, 1973,NumericalTaxonomy. The Principles and Practice of NumericalClassification., San Francisco, CA: W. H. Freeman.
- Sober, Elliott, 1980, “Evolution, Population Thinking, andEssentialism”,Philosophy of Science, 47(3):350–383. doi:10.1086/288942
- –––, 1985, “A Likelihood Justification ofParsimony”,Cladistics, 1(3): 209–233.doi:10.1111/j.1096-0031.1985.tb00424.x
- –––, 1988a, “Likelihood andConvergence”,Philosophy of Science, 55(2):228–237. doi:10.1086/289429
- –––, 1988b,Reconstructing the Past:Parsimony, Evolution, and Inference, Cambridge, MA: MITPress.
- –––, 2004, “The Contest Between Parsimonyand Likelihood”,Systematic Biology, 53(4):644–653. doi:10.1080/10635150490468657
- –––, 2009, “Did Darwin Write the OriginBackwards?”,Proceedings of the National Academy ofSciences, 106(Supplement 1): 10048–10055.doi:10.1073/pnas.0901109106
- –––, 2010,Did Darwin Write the“Origin” Backwards? Philosophical Essays on Darwin’sTheory, Amherst, NY: Prometheus Books.
- –––, 2015,Ockham’s Razors: AUser’s Manual, Cambridge: Cambridge University Press.doi:10.1017/CBO9781107705937
- Sokal, Robert R. and Peter H. A. Sneath, 1963,Principles ofNumerical Taxonomy, San Francisco, CA: W. H. Freeman.
- Steel, Mike and David Penny, 2000, “Parsimony, Likelihood,and the Role of Models in Molecular Phylogenetics”,Molecular Biology and Evolution, 17(6): 839–850.doi:10.1093/oxfordjournals.molbev.a026364
- Sterelny, Kim and Paul Griffiths, 1999,Sex and Death: AnIntroduction to Philosophy of Biology, (Science and ItsConceptual Foundations), Chicago, IL: University of ChicagoPress.
- Sterner, Beckett and Scott Lidgard, 2014, “The NormativeStructure of Mathematization in Systematic Biology”,Studiesin History and Philosophy of Science Part C: Studies in History andPhilosophy of Biological and Biomedical Sciences, 46:44–54. doi:http://dx.doi.org/10.1016/j.shpsc.2014.03.001
- –––, 2018, “Moving Past the SystematicsWars”,Journal of the History of Biology, 51(1):31–67. doi:10.1007/s10739-017-9471-1
- Stevens, Peter F., 2000, “On Characters and CharacterStates: Do Overlapping and Non-Overlapping Variation, Morphology andMolecules All Yield Data of the Same Value?”, in Scotland andPennington 2000: 81–105.
- Sukumaran, Jeet and L. Lacey Knowles, 2017, “MultispeciesCoalescent Delimits Structure, Not Species”,Proceedings ofthe National Academy of Sciences, 114(7): 1607–1612.doi:10.1073/pnas.1607921114
- Sullivan, Jack and Paul Joyce, 2005, “Model Selection inPhylogenetics”,Annual Review of Ecology, Evolution, andSystematics, 36: 445–466.doi:10.1146/annurev.ecolsys.36.102003.152633
- Swofford, David L. and Jack Sullivan, 2009, “PhylogenyInference Based on Parsimony and Other Methods Using PAUP”, inThe Phylogenetic Handbook: A Practical Approach to PhylogeneticAnalysis and Hypothesis Testing, P. Lemey, M. Salemi, and A-M,Vandamme (eds.), Cambridge: Cambridge University Press., pp.267-312.
- Swofford, David L., Peter J. Waddell, John P. Huelsenbeck, PeterG. Foster, Paul O. Lewis, and James S. Rogers, 2001, “Bias inPhylogenetic Estimation and Its Relevance to the Choice betweenParsimony and Likelihood Methods”,Systematic Biology,50(4): 525–539. doi:10.1080/106351501750435086
- Tajima, Fumio, 1983, “Evolutionary Relationship of DNASequences in Finite Populations”,Genetics, 105(2):437–460. doi:10.1093/genetics/105.2.437
- Takezaki, Naoko and Hidenori Nishihara, 2017, “Support forLungfish as the Closest Relative of Tetrapods by Using Slowly EvolvingRay-Finned Fish as the Outgroup”,Genome Biology andEvolution, 9(1): 93–101. doi:10.1093/gbe/evw288
- Tamura, K. and M. Nei, 1993, “Estimation of the Number ofNucleotide Substitutions in the Control Region of Mitochondrial DNA inHumans and Chimpanzees”,Molecular Biology andEvolution, 10(3): 512–526.doi:10.1093/oxfordjournals.molbev.a040023
- Thomson, Keith Stewart, 1991,Living Fossil: The Story of theCoelacanth, New York: W.W. Norton.
- Tuffley, Chris and Mike Steel, 1997, “Links between MaximumLikelihood and Maximum Parsimony under a Simple Model of SiteSubstitution”,Bulletin of Mathematical Biology, 59(3):581–607. doi:10.1007/BF02459467
- Turner, Derek, 2007,Making Prehistory: Historical Science andthe Scientific Realism Debate, Cambridge: Cambridge UniversityPress. doi:10.1017/CBO9780511487385
- van Fraassen, Bas C., 1989,Laws and Symmetry, Oxford:Oxford University Press. doi:10.1093/0198248601.001.0001
- Vassend, Olav B., 2020, “A Verisimilitude Framework forInductive Inference, with an Application to Phylogenetics”,The British Journal for the Philosophy of Science, 71(4):1359–1383. doi:10.1093/bjps/axy054
- Velasco, Joel D., 2007, “Why Non-Uniform Priors on CladesAre Both Unavoidable and Unobjectionable”,MolecularPhylogenetics and Evolution, 45(2): 748–749.doi:10.1016/j.ympev.2007.08.003
- –––, 2008a, “Species Concepts Should NotConflict with Evolutionary History, but Often Do”,Studiesin History and Philosophy of Science Part C: Studies in History andPhilosophy of Biological and Biomedical Sciences, 39(4):407–414. doi:10.1016/j.shpsc.2008.09.007
- –––, 2008b, “The Prior Probabilities ofPhylogenetic Trees”,Biology & Philosophy, 23(4):455–473. doi:10.1007/s10539-007-9105-7
- –––, 2010, “Species, Genes, and the Treeof Life”,The British Journal for the Philosophy ofScience, 61(3): 599–619. doi:10.1093/bjps/axp051
- –––, 2012, “The Future of Systematics:Tree Thinking without the Tree”,Philosophy of Science,79(5): 624–636. doi:10.1086/667878
- –––, 2013, “The Tree of Life”, inThe Cambridge Encyclopedia of Darwin and EvolutionaryThought, Michael Ruse (ed.), Cambridge: Cambridge UniversityPress, 340–345. doi:10.1017/CBO9781139026895.043
- –––, 2019, “The Foundations of ConcordanceViews of Phylogeny”,Philosophy, Theory, and Practice inBiology, 11(July): art. 020.doi:10.3998/ptpbio.16039257.0011.020
- Wagner, Günter P. (ed.), 2000,The Character Concept inEvolutionary Biology, San Diego, CA: Academic Press.doi:10.1016/B978-0-12-730055-9.X5005-8
- Wald, Abraham, 1949, “Note on the Consistency of the MaximumLikelihood Estimate”,The Annals of MathematicalStatistics, 20(4): 595–601.doi:10.1214/aoms/1177729952
- Wang, Lusheng and Tao Jiang, 1994, “On the Complexity ofMultiple Sequence Alignment”,Journal of ComputationalBiology, 1(4): 337–348. doi:10.1089/cmb.1994.1.337
- Weisberg, Michael, 2007, “Three Kinds ofIdealization”,Journal of Philosophy, 104(12):639–659. doi:10.5840/jphil20071041240
- Wheeler, Ward C., Lone Aagesen, Claudia P. Arango, JuliánFaivovich, Taran Grant, Cyrille D’Haese, Daniel Janies, WilliamLeo Smith, Andrés Varón, and Gonzalo Giribet, 2006,Dynamic Homology and Phylogenetic Systematics: A Unified ApproachUsing POY, Washington, DC: American Museum of NaturalHistory.
- Whiting, Michael F., 1998, “Long-Branch Distraction and theStrepsiptera”,Systematic Biology, 47(1):134–137. doi:10.1080/106351598261076
- Whittall, Justen B. and Scott A. Hodges, 2007, “PollinatorShifts Drive Increasingly Long Nectar Spurs in ColumbineFlowers”,Nature, 447(7145): 706–709.doi:10.1038/nature05857
- Wiens, John J., 2004, “The Role of Morphological Data inPhylogeny Reconstruction”,Systematic Biology, 53(4):653–661. doi:10.1080/10635150490472959
- Wiley, E. O., 1975, “Karl R. Popper, Systematics, andClassification: A Reply to Walter Bock and Other EvolutionaryTaxonomists”,Systematic Zoology, 24(2): 233–243.doi:10.1093/sysbio/24.2.233
- –––, 1981,Phylogenetics: The Theory andPractice of Phylogenetic Systematics, New York: Wiley. Secondedition is Wiley and Lieberman 2011.
- Wiley, E. O. and Bruce S. Lieberman, 2011,Phylogenetics:Theory and Practice of Phylogenetic Systematics, second edition,Hoboken, NJ: John Wiley & Sons. See Wiley 1981 for first edition.doi:10.1002/9781118017883
- Williams, David M. and Malte C. Ebach, 2018, “A Cladist Is aSystematist Who Seeks a Natural Classification: Some Comments on Quinn(2017)”,Biology & Philosophy, 33(1–2): art.10. doi:10.1007/s10539-018-9621-7
- Williams, David M. and Malte C. Ebach,Cladistics: A Guide toBiological Classification, 3rd Edition, Cambridge: CambridgeUniversity Press. doi:10.1017/9781139047678.
- Wilson, Robert A., Matthew J. Barker, and Ingo Brigandt, 2007,“When Traditional Essentialism Fails: Biological NaturalKinds”,Philosophical Topics, 35(1–2):189–215. doi:10.5840/philtopics2007351/29
- Wimsatt, William C., 2007,Re-Engineering Philosophy forLimited Beings: Piecewise Approximations to Reality, Cambridge,MA: Harvard University Press.
- Winsor, Mary Pickard, 1976,Starfish, Jellyfish, and the Orderof Life: Issues in Nineteenth Century Science, New Haven, CT:Yale University Press.
- –––, 1995, “The English Debate on Taxonomyand Phylogeny, 1937–1940”,History and Philosophy of theLife Sciences, 17(2): 227–252.
- –––, 2009, “Taxonomy Was the Foundation ofDarwin’s Evolution”,TAXON, 58(1): 43–49.doi:10.1002/tax.581007
- Winther, Rasmus Grønfeldt, 2009, “Character Analysisin Cladistics: Abstraction, Reification, and the Search forObjectivity”,Acta Biotheoretica, 57(1–2):129–162. doi:10.1007/s10441-008-9064-7
- Woese, Carl R. and George E. Fox, 1977, “PhylogeneticStructure of the Prokaryotic Domain: The Primary Kingdoms”,Proceedings of the National Academy of Sciences, 74(11):5088–5090. doi:10.1073/pnas.74.11.5088
- Woese, Carl R., George E. Fox, Lawrence Zablen, Tsuneko Uchida,Linda Bonen, Kenneth Pechman, Bobby J. Lewis, and David Stahl, 1975,“Conservation of Primary Structure in 16S Ribosomal RNA”,Nature, 254(5495): 83–86. doi:10.1038/254083a0
- Woese, Carl R., Otto Kandler, and Mark L. Wheelis, 1990,“Towards a Natural System of Organisms: Proposal for the DomainsArchaea, Bacteria, and Eucarya”,Proceedings of the NationalAcademy of Sciences, 87(12): 4576–4579.doi:10.1073/pnas.87.12.4576
- Yang, Ziheng, 1994, “Statistical Properties of the MaximumLikelihood Method of Phylogenetic Estimation and Comparison withDistance Matrix Methods”,Systematic Biology, 43(3):329–342. doi:10.1093/sysbio/43.3.329
- –––, 1996, “Phylogenetic Analysis UsingParsimony and Likelihood Methods”,Journal of MolecularEvolution, 42(2): 294–307. doi:10.1007/BF02198856
- Yang, Ziheng and Bruce Rannala, 1997, “Bayesian PhylogeneticInference Using DNA Sequences: A Markov Chain Monte CarloMethod”,Molecular Biology and Evolution, 14(7):717–724. doi:10.1093/oxfordjournals.molbev.a025811
- –––, 2010, “Bayesian Species DelimitationUsing Multilocus Sequence Data”,Proceedings of the NationalAcademy of Sciences, 107(20): 9264–9269.doi:10.1073/pnas.0913022107
- –––, 2012, “Molecular Phylogenetics:Principles and Practice”,Nature Reviews Genetics,13(5): 303–314. doi:10.1038/nrg3186
- Zuckerkandl, Emile and Linus Pauling, 1962, “MolecularDisease, Evolution, and Genic Heterogeneity”, inHorizons inBiochemistry: Albert Szent-Györgyi Dedicatory Volume,Michael Kasha and Bernard Pullman (eds.), New York: Academic Press,189–225.
- –––, 1965, “Evolutionary Divergence andConvergence in Proteins”, inEvolving Genes andProteins, Vernon Bryson and Henry J. Vogel (eds.), New York:Academic Press, 97–166.doi:10.1016/B978-1-4832-2734-4.50017-6
- Zuckerkandl, Emile and Walter A. Schroeder, 1961,“Amino-Acid Composition of the Polypeptide Chains of GorillaHaemoglobin”,Nature, 192(4806): 984–985.doi:10.1038/192984a0
Academic Tools
How to cite this entry. Preview the PDF version of this entry at theFriends of the SEP Society. Look up topics and thinkers related to this entry at the Internet Philosophy Ontology Project (InPhO). Enhanced bibliography for this entryatPhilPapers, with links to its database.
Other Internet Resources
- Haber, Matthew H., 2016b, “Transformation, Persistence, and Identity”,Extinct: The Philosophy of Paleontology Blog.
- “Understanding Evolution”, University of California/Berkeley.
- PhyloCode
- PhyloCode Literature, foundational literature, critiques, reply to critics, etc.
- The Society of Systematic Biologists
- “Bears, Species, and DNA”, the University of Utah Genetic Science Learning Center.
Related Entries
adaptationism |biology: philosophy of |developmental biology |developmental biology: evolution and development |empiricism: logical |evolution |evolution: cultural |genetics: ecological |genomics and postgenomics |microbiology, philosophy of |molecular biology |Popper, Karl |species |statistics, philosophy of |underdetermination, of scientific theories
About
Mirror Sites
View this site from another server:
The Stanford Encyclopedia of Philosophy iscopyright © 2024 byThe Metaphysics Research Lab, Department of Philosophy, Stanford University
Library of Congress Catalog Data: ISSN 1095-5054


